Planet Python
Last update: June 08, 2026 04:45 PM UTC
June 08, 2026
Real Python
Python 3.15 Hits Feature Freeze and Other News for June 2026
While the Northern Hemisphere warms up for summer, Python 3.15 went the other way with its beta 1 feature freeze 🥶. Since May 7, the list of what will be included in the next release is final. That list includes a brand-new sentinel built-in that finally standardizes a pattern Python developers have been hand-rolling for decades.
And while AI kept writing code, buggy or not, developers also directed it to look for bugs in code that had been sitting untouched for years. The results were hundreds of bug fixes in Python’s C extensions and in Firefox. Meanwhile, in a quieter corner of the ecosystem, Pydantic forked httpx, kicking off one of the more interesting governance stories of the year.
Time to dig into the Python news from the past month!
Join Now: Click here to join the Real Python Newsletter and you’ll never miss another Python tutorial, course, or news update.
Python Releases and PEP Highlights
The 3.15 release of CPython crossed from alpha into beta, which means its feature set is now frozen, and the Steering Council cleared out a backlog of proposals before the gate closed. Two of those changes will touch the code you write every day.
Beta 1 Marks the 3.15 Feature Freeze
Last month, the eighth and final alpha rolled out as the runway to the beta phase. With Python 3.15.0b1 on May 7 came the feature freeze, which means that from here until the final release of 3.15, the core team works only on bug fixes and polishing.
That makes the beta releases a good moment to step back and look at the headline features of 3.15, which are now locked:
- Explicit lazy imports (PEP 810) for faster startup
- A
frozendictbuilt-in (PEP 814) for immutable mappings - A
sentinelbuilt-in (PEP 661), which you’ll dig into below - Unpacking in comprehensions (PEP 798)
- UTF-8 as the default encoding (PEP 686)
- A stable ABI for free-threaded builds (PEP 803), plus C-API modernization (PEPs 820 and 793) that should make it easier to write C extensions that work across Python versions
- A new sampling profiler in the standard library (PEP 799) for low-overhead profiling
The JIT compiler also gets faster, with the beta announcement citing an 8–9 percent geometric-mean improvement on x86-64 Linux. If you’ve been putting off testing your code against 3.15, then now is the time to get started! The API surface won’t shift under you anymore, and your feedback will help catch regressions before the release candidate phase.
Note: Beta builds are for testing, not production. Install the pre-release version, run your test suite against 3.15, and report anything that breaks while there’s still time to fix it before the release candidate.
The first round of improvements already landed with beta 2 on June 2, and the next big checkpoint is the release candidate phase on August 4, with the final release expected, as usual, this fall.
A Built-in sentinel Lands in Python 3.15
Here’s the new feature that you’ll likely want to reach for. If you’ve ever needed to tell the difference between a caller passing None and a caller passing nothing at all, then you’ve probably written something like this:
_MISSING = object()
def update(value=_MISSING):
if value is _MISSING:
... # No value was provided
It works, but it has rough edges. The repr() is an unhelpful <object object at 0x7f...>, the marker can’t be used cleanly in type annotations, and its identity doesn’t survive copying or pickling. PEP 661 replaces the idiom with a new sentinel built-in:
MISSING = sentinel("MISSING")
def update(value: int | MISSING = MISSING) -> None:
if value is MISSING:
... # No value was provided
The signature is sentinel(name, /, *, repr=None), and the result is a unique truthy object whose default repr() is the name you gave it, so MISSING shows up as MISSING in tracebacks instead of a memory address.
Note: Sentinels and None solve related but different problems. If you’re still fuzzy on when None is the right tool, then Real Python’s guide to Python’s None is worth revisiting.
Because the sentinel is its own type, you can drop it straight into annotations like int | MISSING without reaching for Literal. The PEP was first submitted back in 2021, so it’s satisfying to see it cross the finish line.
PEP 829 Graduates From Draft to Accepted
Last month’s roundup featured PEP 829 while it was still a draft. It’s since been accepted for Python 3.15, so the change is now official.
As a quick recap, .pth files in your site-packages directory can do two things:
- Extend
sys.path - Run arbitrary code through
importlines that Python feeds directly toexec()at startup
Read the full article at https://realpython.com/python-news-june-2026/ »
[ Improve Your Python With 🐍 Python Tricks 💌 – Get a short & sweet Python Trick delivered to your inbox every couple of days. >> Click here to learn more and see examples ]
death and gravity
Ordered key sharding in DynamoDB
So, you want to keep a sorted index in DynamoDB, but for whatever reason – usually throughput-related – it won't fit on a single partition. What do you do?
Today, we look at the available solutions, do the math, and find out which is best.
Tip
This worked example is part of my DynamoDB crash course series.
Contents- Requirements
- A sparse index is almost enough
- But scan results are not ordered
- But a single partition key causes throttling
- But random suffixes are random
- But hash suffixes are not ordered
- But there are a lot of first characters
- But some first bytes need multiple shards
- But tries and prefix ranges are complicated
- But the prefix distribution can change
Requirements #
Say you're using single table design with a table of artists, albums, and songs.1
You keep an artist's items in a single collection
(aka same partition key),
and use sort keys artist, album#{Album}, and song#{Album}#{Song},
depending on their type:
# table Music (partition key: Artist, sort key: sk)
Solar Fields: !btree
'album#Leaving Home': { Album: Leaving Home, ... }
'artist': { ... }
'song#Leaving Home#Air Song': { ... }
'song#Leaving Home#Monogram': { ... }
To list albums without doing a full table scan, you need a global secondary index.
Let's come up with some reasonable requirements; the GSI should support:
- items up 500 bytes (we project additional attributes besides the keys)
- 10,000 queries/second, max 100 items/query, sorted alphabetically
- list all albums
- list albums by title
- 10,000 writes/second (to avoid write throttling during imports)
A sparse index is almost enough #
One way to do it is to use a dedicated sparse index, taking advantage of the fact that items with missing index keys don't appear in the index.
If only albums have an Album attribute, we just create a new GSI:
# GSI Albums (partition key: Album, sort key: Artist)
Leaving Home: !btree
International Pony: { sk: 'album#Leaving Home', ... }
Solar Fields: { sk: 'album#Leaving Home', ... }
Heavy Migration: !btree
Dday One: { sk: 'album#Heavy Migration', ...}
If songs have an Album too, we add a dedicated AlbumsPK attribute instead.
In many ways, this is the ideal solution. To list all albums, we scan the index. To list albums by title, we query an index partition key. We have lots of unique partition keys with items spread pretty evenly across them, which should prevent throttling.
But scan results are not ordered #
...except scan results are not ordered, so we're missing the sorted alphabetically part.
What is ordered are sort keys, so we can use a single index collection instead:
# GSI GSI1 (partition key: gsi1pk, sort key: gsi1sk)
'albums': !btree
Heavy Migration: { Artist: Dday One, sk: 'album#Heavy Migration', ... }
Leaving Home: { Artist: Solar Fields, sk: 'album#Leaving Home', ... }
Leaving Home: { Artist: International Pony, sk: 'album#Leaving Home', ... }
This is also seemingly ideal. To list all albums, we query the entire index partition key. To list albums by title, we use a sort key. The results are sorted as required, and there's no limit on the number of items in a collection.
But a single partition key causes throttling #
However, there are per-partition limits of 24 MB/s for reads and 1 MB/s for writes.
Let's see how they compare to our requirements:
- reads: 500 bytes/item * 10k queries/s * 100 items/query = 500 MB/s (~21x)
- writes: 500 bytes/item * 10k items/s = 5 MB/s (5x)
Uh-oh, turns out we need 21 times the throughput one partition can deliver.
One way to spread the load is
sharding,
using multiple synthetic partition keys of the form album#{shard_id}.
A common option for the shard id is a random number from a known range,
e.g. album#{randrange(21)}:
# GSI GSI1 (partition key: gsi1pk, sort key: gsi1sk)
'album#1': !btree
Leaving Home: { Artist: Solar Fields, ... }
'album#12': !btree
Heavy Migration: { Artist: Dday One, ... }
'album#20': !btree
Leaving Home: { Artist: International Pony, ... }
To list all albums, query each shard in turn:
for shard in range(21):
for item in dynamodb.query(f"album#{shard}"):
yield item
But random suffixes are random #
There's a problem, though – with random shard ids we can't easily list albums by title, since albums with the same title may end up on any shard.
A better option is to calculate the shard id from the album title using a hash function:
def hash(s):
return int.from_bytes(sha256(s.encode()).digest())
def album_shard_id(album_title):
return hash(album_title) % 21
# GSI GSI1 (partition key: gsi1pk, sort key: gsi1sk)
'album#6': !btree
Leaving Home: { Artist: Solar Fields, ... }
Leaving Home: { Artist: International Pony, ... }
'album#8': !btree
Heavy Migration: { Artist: Dday One, ... }
To list albums by title:
dynamodb.query(f"album#{album_shard_id(album_title)}", sk=album_title, index='GSI1')
But hash suffixes are not ordered #
That takes care of throughput, but now results aren't sorted alphabetically anymore. We can sort items within each shard using the sort key, but they are spread uniformly across shards, and there's no order between shards.
Maybe we could use the first letter as shard id instead?
Of course, we have to account for some first letters being more frequent than others. In this case, we can approximate the actual distribution by using MusicBrainz data.
There are 5.5 million albums:
>>> import polars as pl
>>> titles = pl.read_csv(
... 'mbdump/release',
... has_header=False,
... separator='\t',
... quote_char=None,
... columns=[2],
... new_columns=['title'],
... )[:,0]
>>> titles.count()
5535986
...but only 3.3 million unique titles, partly due to different releases of the same album, partly due to some titles being more popular – a few of them, really popular:
>>> titles.value_counts(sort=True)
shape: (3_370_505, 2)
title count
Greatest Hits 4638
Demo 3140
… …
Salsa salsa 1
Glamour: Deluxe 1
>>> titles.unique_counts().quantile([.9, .99, .999, .9999, 1])
[2.0, 11.0, 58.0, 221.0, 4638.0]
Let's look at first characters:
>>> normalized = titles.str.to_lowercase().str.normalize('NFKD').sort()
>>> normalized.str.slice(0, 1).value_counts(sort=True)
shape: (5_402, 2)
title count
t 584760
s 509065
a 317513
l 298757
… …
🫀 1
🫂 1
🫧 1
1
But there are a lot of first characters #
5402?! Indeed, there's more to Unicode than the Latin alphabet:
>>> normalized.str.slice(0, 1).unique().sample(10).to_list()
['學', '舒', 'і', '进', '੦', '潮', '向', '妳', '陳', '🍅']
And it's actually worse than that – there are five thousand characters in our dataset, but there are hundreds of thousands of possible Unicode characters.
This is not a problem when adding the albums, but it is a problem when listing them, since we need to enumerate all the shards in a reasonable amount of time (and most shards being empty doesn't help, either).
As a very bad compromise, we could use the first byte of the UTF-8 encoding instead; this caps the number of shard ids at 256, and at least Latin titles would be sorted (I did say it's a bad compromise). There:
>>> firstbytes = normalized.map_elements(str.encode).bin.slice(0, 1)
>>> firstbytes.value_counts(sort=True)
shape: (136, 2)
title count
b"t" 584760
b"s" 509065
b"a" 317513
b"l" 298757
… …
b"\xd4" 2
b"U" 1
b"\xee" 1
b"\xf3" 1
But some first bytes need multiple shards #
We knew the first byte distribution would be skewed, but some of them don't even fit on a single shard (and it gets worse the more shards we need):
>>> shard_count = 21
>>> firstbytes.value_counts(sort=True).with_columns(
... pl.col('count') / (len(firstbytes) / shard_count)
... ).head()
shape: (5, 2)
title count
b"t" 2.218206
b"s" 1.931068
b"a" 1.204442
b"l" 1.133294
b"b" 1.106949
We're back to where we started: how do we sort between shards with the same prefix?
We don't – we find longer prefixes that fit in one shard.
That sounds like the perfect job for a trie (aka prefix tree). This would also allow us to switch back to characters, and merge small prefixes into ranges until each range fits one shard. But that's complicated, and as often the case, there must be a better way.2
But tries and prefix ranges are complicated #
We're looking for contiguous ranges, each of a certain size. Tries are good for finding the shortest prefix, but we don't really care about prefix length.
Why not just split the sorted titles into N equal ranges instead? This takes care of the uneven distribution:
>>> boundaries = normalized.gather_every(2000).str.slice(0, 4)
>>> boundaries.value_counts(sort=True)
shape: (2_103, 2)
title count
the 161
live 23
… …
風吹けは 1
魔法少女 1
...provided a long enough prefix:
>>> boundaries = normalized.gather_every(2000).str.slice(0, 16)
>>> boundaries.value_counts(sort=True).filter(pl.col('count') > 1)
shape: (3, 2)
title count
greatest hits 3
demo 2
the very best of 2
...almost there:
>>> boundaries = normalized.gather_every(2000).str.slice(0, 20)
>>> boundaries.value_counts(sort=True).filter(pl.col('count') > 1)
shape: (2, 2)
title count
greatest hits 3
demo 2
This highlights another problem – if the shard size is too small, there may be more than a shard's worth of albums with identical titles; we can fix this by using another, random suffix (ordering doesn't matter anymore, since they have the same title).
Thankfully, our shards are huge, so it's not an issue:
>>> shard_size = int(math.ceil(len(normalized) / shard_count))
>>> shard_size
263619
>>> boundaries = normalized.gather_every(shard_size).str.slice(0, 20)
To use this, save the list of boundaries in code, and find the index of the biggest boundary smaller than a given album title:
import bisect
import unicodedata
ALBUM_TITLE_BOUNDARIES = [
'', # replaced with the smallest possible string
'agartha',
'barstow / crazy',
'can you feel it',
'cyan rot',
'dreams take over eve',
'feud semiotics (rb. ',
'grave poetry',
'i live',
'kannaval',
'live in florence',
"mir ist's gleich / i",
'notice',
'platforms ep',
'rituals',
'skylten',
'surtr / absorbed',
'the human touch',
'tonttujen jouluyö: ',
'walking away',
'голос',
]
def album_shard_id(album_title):
normalized = unicodedata.normalize('NFKD', album_title.lower())
return bisect.bisect(ALBUM_TITLE_BOUNDARIES, normalized) - 1
>>> album_shard_id('2 Pie Island')
0
>>> album_shard_id('Heavy Migration')
7
>>> album_shard_id('Leaving Home')
9
>>> album_shard_id('Space Cadet')
15
But the prefix distribution can change #
We were lucky to have data on the prefix distribution, but that's not always the case, and even if it is, the distribution can change over time.
For example, the last of the 21 shards above starts in the Cyrillic Unicode block, which means most existing scripts go into a single shard. What if we import 1 million Chinese and Japanese albums at some point?
One way to deal with this is to give more weight to known gaps in the data. Another is to have more shards from the start to account for unknown gaps – 210 shards instead of 21 sounds pretty reasonable.
If all else fails, you can move to a new index with more shards, but that comes with its own complications, so it's best to get it roughly right from the start.
Anyway, that's it for now.
Learned something new today? Share it with others, it really helps!
Want to know when new articles come out? Subscribe here to get new stuff straight to your inbox!
This is a simplified example; as the MusicBrainz database shows, the schema for this kind of thing would be way more complicated in practice. [return]
You're welcome to try, though, especially if you're preparing for an interview. [return]
Wingware
Wing Python IDE 12 Early Access - June 8, 2026
Wing 12 is now available as an early access release that focuses on AI agent driven development. Wing 12 introduces deep integration with Claude Code, including a dedicated Claude Code tool, a new Tasks tool for planning, executing, and reviewing AI agent work, and a set of MCP servers that allow agents to work more efficiently by giving them access to Wing's source code analysis, unit testing, and debugger features.
For those using Claude Code, Wing 12 broadens the focus from code-centric direct development to also include managing multiple concurrent AI agents. Of course all of Wing's classic IDE features are still available -- the powerful debugger, deep code analysis and warnings, full-featured editor, project & package management, and much more.
Wing 12 also adds true pseudo-terminal support to OS Commands and Debug I/O, reenvisions the OS Commands tool so that commands can be dragged to tool or editor splits, reorganizes the Tools menu, adds search and back/forward navigation to the Preferences dialog, supports automatic test discovery, speeds up detection of externally modified files, and makes many other improvements.
For more information, see the Wingware Early Access Program. Anyone can participate just by downloading the release.
If you have questions, please don't hesitate to contact us at support@wingware.com.
June 07, 2026
Eli Bendersky
Plugins case study: mdBook preprocessors
mdBook is a tool for easily creating books out of Markdown files. It's very popular in the Rust ecosystem, where it's used (among other things) to publish the official Rust book.
mdBook has a simple yet effective plugin mechanism that can be used to modify the book output in arbitrary ways, using any programming language or tool. This post describes the mechanism and how it aligns with the fundamental concepts of plugin infrastructures.
mdBook preprocessors
mdBook's architecture is pretty simple: your contents go into a directory tree of Markdown files. mdBook then renders these into a book, with one file per chapter. The book's output is HTML by default, but mdBook supports other outputs like PDF.
The preprocessor mechanism lets us register an arbitrary program that runs on the book's source after it's loaded from Markdown files; this program can modify the book's contents in any way it wishes before it all gets sent to the renderer for generating output.
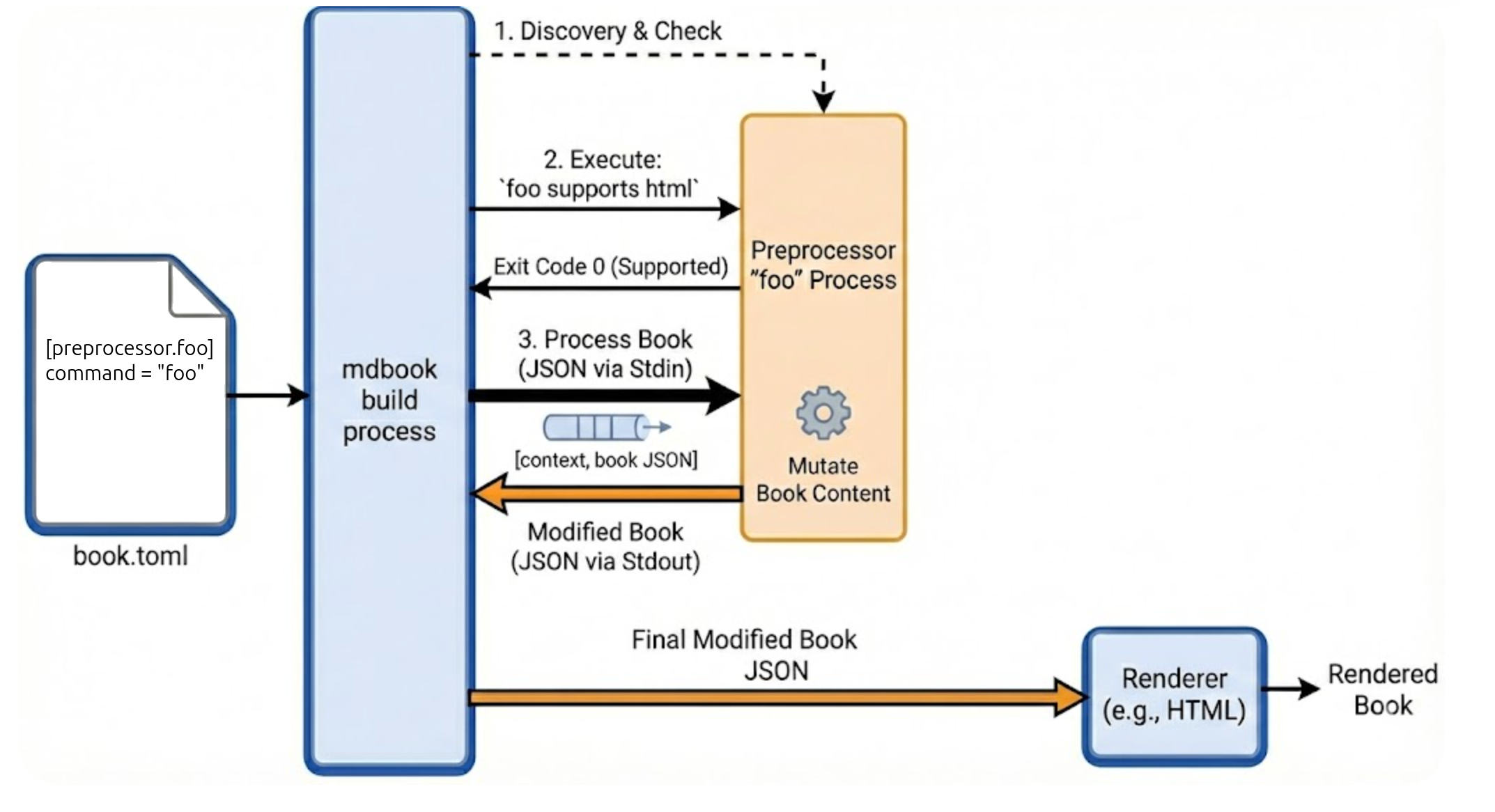
The official documentation explains this process very well.
Sample plugin
I rewrote my classical "nacrissist" plugin for mdBook; the code is available here.
In fact, there are two renditions of the same plugin there:
- One in Python, to demonstrate how mdBook can invoke preprocessors written in any programming language.
- One in Rust, to demonstrate how mdBook exposes an application API to plugins written in Rust (since mdBook is itself written in Rust).
Fundamental plugin concepts in this case study
Let's see how this case study of mdBook preprocessors measures against the Fundamental plugin concepts that were covered several times on this blog.
Discovery
Discovery in mdBook is very explicit. For every plugin we want mdBook to use, it has to be listed in the project's book.toml configuration file. For example, in the code sample for this post, the Python narcissist plugin is noted in book.toml as follows:
[preprocessor.narcissistpy]
command = "python3 ../preprocessor-python-narcissist/narcissist.py"
Each preprocessor is a command for mdBook to execute in a sub-process. Here it uses Python, but it can be anything else that can be validly executed.
Registration
For the purpose of registration, mdBook actually invokes the plugin command twice. The first time, it passes the arguments supports <renderer> where <renderer> is the name of the renderer (e.g. html). If the command returns 0, it means the preprocessor supports this renderer; otherwise, it doesn't.
In the second invocation, mdBook passes some metadata plus the entire book in JSON format to the preprocessor through stdin, and expects the preprocessor to return the modified book as JSON to stdout (using the same schema).
Hooks
In terms of hooks, mdBook takes a very coarse-grained approach. The preprocessor gets the entire book in a single JSON object (along with a context object that contains metadata), and is expected to emit the entire modified book in a single JSON object. It's up to the preprocessor to figure out which parts of the book to read and which parts to modify.
Given that books and other documentation typically have limited sizes, this is a reasonable design choice. Even tens of MiB of JSON-encoded data are very quick to pass between sub-processes via stdout and marshal/unmarshal. But we wouldn't be able to implement Wikipedia using this design.
Exposing an application API to plugins
This is tricky, given that the preprocessor mechanism is language-agnostic. Here, mdBook only offers additional utilities to preprocessors implemented in Rust. These get access to mdBook's API to unmarshal the JSON representing the context metadata and book's contents. mdBook offers the Preprocessor trait Rust preprocessors can implement, which makes it easier to wrangle the book's contents. See my Rust version of the narcissist preprocessor for a basic example of this.
Renderers / backends
Actually, mdBook has another plugin mechanism, but it's very similar conceptually to preprocessors. A renderer (also called a backend in some of mdBook's own doc pages) takes the same input as a preprocessor, but is free to do whatever it wants with it. The default renderer emits the HTML for the book; other renderers can do other things.
The idea is that the book can go through multiple preprocessors, but at the end a single renderer.
The data a renderer receives is exactly the same as a preprocessor - JSON encoded book contents. Due to this similarity, there's no real point getting deeper into renderers in this post.
June 06, 2026
Armin Ronacher
Communities of Not
There is a strange thing that happens in communities that gather around abstinence from something: identity from opposition. At their best these communities are not just negative: childfree spaces can be about autonomy, choice and acceptance, anti-car spaces about safer streets and transit, and LLM-skeptical developer spaces about the future of labor, code quality and slop1. But the thing being refused often does not go away and instead becomes the main subject of the community’s identity.
That would be fine if it stayed at criticism, maybe even angry criticism, but more often than not it turns into policing and hatred towards others. An influencer without children becomes a parent, an urban bike commuter by choice buys a Porsche, a respected developer tries LLMs, and the community feels betrayed because it assumed they were members of the same tribe. The expulsion of that person (who never signed up to be a community member) is entirely imaginary but the punishment that the community unleashes is not: people pile on and shame them, quote them out of context and turn their weakest moments into proof that the person was always unserious, a sharlatan or should not be listened to.
I do not think the answer is to tell people to stop paying attention. Cars shape cities even for people who cycle, children influence politics, workplaces and taxes even for people who do not have them. For us developers, LLMs show up in editors, issue trackers, hiring conversations, management pressure and code reviews whether we asked for them or not. Resisting that can be legitimate but that is no excuse for using one’s rejection to justify shitty mob behavior.
I understand the thinking all too well, because I have done versions of this myself in the past. It took me a while to become more accepting of other people’s worldviews that diverge from mine. Whatever insecurities we have, finding a group of others sharing them can be comforting. The danger is that being part of a crowd of negativity can easily make us part of collective harassment.
I can only encourage you to breathe, slow down, de-escalate when given the chance, and resist the temptation to always assume the most catastrophic reading. Default to being open to new things. Being negative towards something, and making that ones identity, is an easy trap to fall into.
-
These examples are not meant as equivalents. The recent mob against rsync is the LLM version that prompted this post. I picked the others because I’m familiar with those communities and they all show similar cases of personal choices being interpreted as betrayal.↩
June 05, 2026
Kay Hayen
Nuitka Release 4.1
This is to inform you about the new stable release of Nuitka. It is the extremely compatible Python compiler, “download now”.
This release adds many new features and corrections with a focus on async code compatibility, missing generics features, and Python 3.14 compatibility and Python compilation scalability yet again.
Bug Fixes
Python 3.14: Fix, decorators were breaking when disabling deferred annotations. (Fixed in 4.0.1 already.)
Fix, nested loops could have wrong traces lead to mis-optimization. (Fixed in 4.0.1 already.)
Plugins: Fix, run-time check of package configuration was incorrect. (Fixed in 4.0.1 already.)
Compatibility: Fix,
__builtins__lacked necessary compatibility in compiled functions. (Fixed in 4.0.1 already.)Distutils: Fix, incorrect UTF-8 decoding was used for TOML input file parsing. (Fixed in 4.0.1 already.)
Fix, multiple hard value assignments could cause compile time crashes. (Fixed in 4.0.1 already.)
Fix, string concatenation was not properly annotating exception exits. (Fixed in 4.0.2 already.)
Windows: Fix,
--verbose-outputand--show-modules-outputdid not work with forward slashes. (Fixed in 4.0.2 already.)Python 3.14: Fix, there were various compatibility issues including dictionary watchers and inline values. (Fixed in 4.0.2 already.)
Python 3.14: Fix, stack pointer initialization to
localspluswas incorrect to avoid garbage collection issues. (Fixed in 4.0.2 already.)Python 3.12+: Fix, generic type variable scoping in classes was incorrect. (Fixed in 4.0.2 already.)
Python 3.12+: Fix, there were various issues with function generics. (Fixed in 4.0.2 already.)
Python 3.8+: Fix, names in named expressions were not mangled. (Fixed in 4.0.2 already.)
Plugins: Fix, module checksums were not robust against quoting style of module-name entry in YAML configurations. (Fixed in 4.0.2 already.)
Plugins: Fix, doing imports in queried expressions caused corruption. (Fixed in 4.0.2 already.)
UI: Fix, support for
uv_buildin the--projectoption was broken. (Fixed in 4.0.2 already.)Compatibility: Fix, names assigned in assignment expressions were not mangled. (Fixed in 4.0.2 already.)
Python 3.12+: Fix, there were still various issues with function generics. (Fixed in 4.0.3 already.)
Clang: Fix, debug mode was disabled for clang generally, but only ClangCL and macOS Clang didn’t want it. (Fixed in 4.0.3 already.)
Zig: Fix,
--windows-console-mode=attach|disablewas not working when using Zig. (Fixed in 4.0.3 already.)macOS: Fix, yet another way self dependencies can look like, needed to have support added. (Fixed in 4.0.3 already.)
Python 3.12+: Fix, generic types in classes had bugs with multiple type variables. (Fixed in 4.0.3 already.)
Scons: Fix, repeated builds were not producing binary identical results. (Fixed in 4.0.3 already.)
Scons: Fix, compiling with newer Python versions did not fall back to Zig when the developer prompt MSVC was unusable, and error reporting could crash. (Fixed in 4.0.4 already.)
Zig: Fix, the workaround for Windows console mode
attachordisablewas incorrectly applied on non-Windows platforms. (Fixed in 4.0.4 already.)Standalone: Fix, linking with Python Build Standalone failed because
libHacl_Hash_SHA2was not filtered out unconditionally. (Fixed in 4.0.4 already.)Python 3.6+: Fix, exceptions like
CancelledErrorthrown into an async generator awaiting an inner awaitable could be swallowed, causing crashes. (Fixed in 4.0.4 already.)Fix, not all ordered set modules accepted generators for update. (Fixed in 4.0.5 already.)
Plugins: Disabled warning about rebuilding the
pytokensextension module. (Fixed in 4.0.5 already.)Standalone: Filtered
libHacl_Hash_SHA2from link libs unconditionally. (Fixed in 4.0.5 already.)Debugging: Disabled unusable unicode consistency checks for Python versions 3.4 to 3.6. (Fixed in 4.0.5 already.)
Python3.12+ Avoided cloning call nodes on class level which caused issues with generic functions in combination with decorators. (Added in 4.0.5 already.)
Python 3.12+: Added support for generic type variables in
async deffunctions. (Added in 4.0.5 already.)UI: Fix, flushing outputs for prompts was not working in all cases when progress bars were enabled. (Fixed in 4.0.6 already.)
UI: Fix, unused variable warnings were missing at C compile time when using
zigas a C compiler. (Fixed in 4.0.6 already.)Scons: Fix, forced stdout and stderr paths as a feature was broken. (Fixed in 4.0.6 already.)
Fix, replacing a branch did not accurately track shared active variables causing optimization crashes. (Fixed in 4.0.7 already.)
macOS: Fix, failed to remove extended attributes because files need to be made writable first. (Fixed in 4.0.7 already.)
Fix, dict
popandsetdefaultusing with:=rewrites lacked exception-exit annotations for un-hashable keys. (Fixed in 4.0.8 already.)Python 3.13: Fix, the
__parameters__attribute of generic classes was not working. (Fixed in 4.0.8 already.)Python 3.11+: Fix, starred arguments were not working as type variables. (Fixed in 4.0.8 already.)
Python2: Fix,
FileNotFoundErrorcompatibility fallback handling was not working properly. (Fixed in 4.0.8 already.)Compatibility: Fix, loop ownership check in value traces was missing, causing issues with nested loops.
Windows: Improved
--windows-console-mode=attachto properly handle console handles, enabling cases likeos.systemto work nicely.Python2: Fix, there was a compatibility issue where providing default values to the
mkdtempfunction was failing.Windows: Fix, there were spurious issues with C23 embedding in 32-bit MinGW64 by switching to
coff_objresource mode for it as well.Plugins: Fix, the
post-import-codeexecution could fail because the triggering sub-package was not yet available insys.modules.UI: Fix, listing package DLLs with
--list-package-dllswas broken due to recent plugin lifecycle changes.UI: Fix,
--list-package-exewas not working properly on non-Windows platforms failing to detect executable files correctly.UI: Handled paths starting with
{PROGRAM_DIR}the same as a relative path when parsing the--onefile-tempdir-specoption.Plugins: Followed multiprocessing
forkserverchanges for newer Python versions.Python 3.12+: Fix, generic class type parameters handling was incorrect.
Python 3.12: Fix, deferred evaluation of type aliases was failing.
Python 3.12+: Aligned
sumbuilt-in float summation with CPython’s compensated sum for better accuracy.Python 3.10+: Fix, uncompiled coroutine
throw()return handling was incorrect, restoring completed coroutine results viaStopIteration.valuerather than exposing them as ordinary return values to the outer await chain.Python 3.13+: Fix, uncompiled coroutine
cancel()/awaitsuspension handling was incorrect, improved to ensure integration compatibility.macOS: Made finding
create-dmgmore robustly by also checking the Homebrew path for Intel and fromPATHproperly.Compatibility: Fix, class frames were not exposing frame locals.
UI: Detected
static-libpythonproblems, which affected some forms of Anaconda.Distutils: Rejected
--projectmixed with--mainarguments as it is not useful.macOS: Fix,
zigfromPATHor fromziglangwas not being used.Distutils: Fix, the wrong
module-rootconfig value was being checked foruvbuild backend.macOS: Fix, was attempting to change removed (rejected) DLLs, which of course failed and errored out.
Python 3.14: Fix, tuple reuse was not fully compatible, potentially causing crashes due to outdated hash caches.
Fix, fake modules were still being attempted to located when imported by other code, which could conflict with existing modules.
Python 3.5+: Fix, failed to send uncompiled coroutines the sent in value in
yield from.Fix, older
gcccompilers lacking newer intrinsic methods had compilation issues that needed to be addressed.Standalone: Fix, multiphase module extension modules with post-load code were not working properly.
Fix, Avoid using the non-inline copy of
pkg_resourceswith the inline copy of Jinja2. These could mismatch and cause errors.Fix, loops could make releasing of previous values very unclear, causing optimization errors.
Fix,
incbinresource mode was not working with oldgccC++ fallback.Python 3.4 to 3.6: Fix, bytecode demotion was not working properly for these versions, also bytecode only files not working.
Plugins: Added a check for the broken
patchelfversions 0.10 and 0.11 to prevent breaking Qt plugins.Android: Allowed
patchelfversion 0.18 on Android.Windows: Fix, the header path for self uninstalled Python was not detected correctly.
Release: Fix, inclusion of the
pkg_resourcesinline copy for Python 2 to source distributions was missing.UI: Detected the OBS versions of SUSE Linux better.
Suse: Allowed using
patchelf0.18.0 there too.Python 3.11: Fix, package and module dicts were not aligned close enough to avoid a CPython bug.
Fix, unbound compiled methods could crash when called without an object passed.
Standalone: Fix, multiphase module extension modules with postload. (Fixed in 4.0.8 already.)
Onefile: Fix, while waiting for the child, it may already be terminated.
macOS: Removed existing absolute rpaths for Homebrew and MacPorts.
Python 3.14: Avoided warning in CPython headers.
Python 3.14: Followed allocator changes more closely.
Compatibility: Avoided using
pkg_resourcesfor Jinja2 template location for loading.No-GIL: Applied some bug fixes to get basic things to work.
Package Support
Standalone: Add support for newer
paddleversion. (Added in 4.0.1 already.)Standalone: Add workaround for refcount checks of
pandas. (Fixed in 4.0.1 already.)Standalone: Add support for newer
h5pyversion. (Added in 4.0.2 already.)Standalone: Add support for newer
scipypackage. (Added in 4.0.2 already.)Plugins: Revert accidental
os.getenvoveros.environ.getchanges in anti-bloat configurations that stopped them from working. Affected packages arenetworkx,persistent, andtensorflow. (Fixed in 4.0.5 already.)Standalone: Added missing DLLs for
openvino. (Added in 4.0.7 already.)Enhanced the package configuration YAML schema by adding the
relative_toparameter forfrom_filenamesDLL specification, avoiding error-prone purely relative paths.Standalone: Fix,
flet_desktopapp assets were missing, now preserving the packaged runtime and sidecar DLLs.Standalone: Added support for the
tyropackage.Standalone: Added data files for the
perfettopackage.Standalone: Added support for
anyioprocess forking.Standalone: Added support for the
plotly.graphpackage.Anaconda: Fix, dependencies for the
numpyconda package on Windows were incorrect.Plugins: Enhanced the auto-icon hack in PySide6 to use compatible class names.
Standalone: Fix, Qt libraries were duplicated with
PySide6WebEngine framework support on macOS.Plugins: Fix, automatic detection of
mypycruntime dependencies was including all top level modules of the containing package by accident. (Fixed in 4.0.5 already.)Anaconda: Fix,
delvewheelplugin was not working with Python 3.8+. This enhances compatibility with installed PyPI packages that use it for their DLLs. (Fixed in 4.0.6 already.)Plugins: Fix, our protection workaround could confuse methods used with
PySide6.
New Features
UI: Added the
--recommended-python-versionoption to display recommended Python versions for supported, working, or commercial usage.UI: Add message to inform users about
Nuitka[onefile]if compression is not installed. (Added in 4.0.1 already.)UI: Add support for
uv_buildin the--projectoption. (Added in 4.0.1 already.)Onefile: Allow extra includes as well. (Added in 4.0.2 already.)
UI: Add
nuitka-project-setfeature to define project variables, checking for collisions with reserved runtime variables. (Added in 4.0.2 already.)Scons: Added new option to select
--reproduciblebuilds or not. (Added in 4.0.6 already.)Python 3.10+: Added support for
importlib.metadata.package_distributions(). (Added in 4.0.8 already.)Plugins: Added support for the multiprocessing
forkservercontext. (Added in 4.0.8 already, for 4.1 Python 3.6 and earlier, as well as 3.14 support were added too.)Reports: Added structured resource usage (
rusage) performance information to compilation reports.Reports: Included individual module-level C compiler caching (
ccache/clcache) statistics in compilation reports.Added support for detecting and correctly resolving the Python prefix for the
PyEnv on HomebrewPython flavor.macOS: Added support for
rusageinformation for Scons.UI: Added the
__compiled__.extension_filenameattribute to give the real filename of the containing extension module.Windows: Added support for
--clangor ARM. (Added in 4.0.8 already.)Windows: Added support for resources names as not just integers, important when we copy them from template files.
MacPorts: Added basic support for this Python flavor. More work will be needed to get it to work fully though.
Optimization
Avoid including
importlib._bootstrapandimportlib._bootstrap_external. (Added in 4.0.1 already.)Linux: Cached the
syscallused for time keeping during compilation to avoid loadinglibcfor each trace. (Added in 4.0.8 already.)UI: Output a warning for modules that remain unfinished after the third optimization pass.
Added an extra micro pass trigger when new variables are introduced or variable usage changes severely, ensuring optimizations are fully propagated, avoiding unnecessary extra full passes.
Provided scripts to compile Python statically with PGO tailored for Nuitka on Linux, Windows, and macOS.
Added support for running the Data Composer tool from a compiled Nuitka binary without spawning an uncompiled Python process.
Enhanced the usage of
vectorcallforPyCFunctionobjects by directly checking for its presence instead of relying purely on flags, allowing more frequent use of this faster execution path.Cached frequently used declarations for top-level variables to speed up C code generation.
Sped up trace collection merging by avoiding unnecessary set creation and using a set instead of a list for escaped traces.
Optimized plugin hook execution by tracking overloaded methods and added an option to show plugin usage statistics.
Improved performance of module location by avoiding unnecessary module name reconstruction and redundant filesystem checks for pre-loaded packages.
Improved the caching of distribution name lookups to effectively avoid repeated IO operations across all package types.
Plugins: Cached callback plugin dispatch for
onFunctionBodyParsingandonClassBodyParsingto skip argument computation when no plugin overrides them.Python 3.13: Handled sub-packages of
pathlibas hard modules.Handled hard attributes through merge traces as well.
Made constant blobs more compact by avoiding repeated identifiers and unnecessary fields.
Enhanced Python compilation scripts further. (Fixed in 4.0.8 already.)
Recognized late incomplete variables better. (Fixed in 4.0.8 already.)
Made constant blobs more compact. (Fixed in 4.0.8 already.)
Optimized calls with only constant keywords and variable posargs too.
Anti-Bloat
Fix, memory bloat occurred when C compiling
sqlalchemy. (Fixed in 4.0.2 already.)Avoid using
pydocinPySimpleGUI. (Added in 4.0.2 already.)Avoided using
doctestfromzodbpickle. (Added in 4.0.5 already.)Avoided inclusion of
cythonwhen usingpyav. (Added in 4.0.7 already.)Avoided including
typing_extensionswhen usingnumpy. (Added in 4.0.7 already.)
Organizational
UI: Relocated the warning about the available source code of extension modules to be evaluated at a more appropriate time.
Debian: Remove recommendation for
libfuse2package as it is no longer useful.Debian: Used
platformdirsinstead ofappdirs.Debugging: Removed Python 3.11+ restriction for
clang-formatas it is available everywhere, even Python 2.7, and we still want nicely formatted code when we read things. (Added in 4.0.6 already.)Removed no longer useful inline copy of
wax_off. We have our own stubs generator project.Release: Added missing package to the CI container for building Nuitka Debian packages.
Developer: Updated AI instructions for creating Minimal Reproducible Examples (MRE) to skip unneeded C compilation.
Debugging: Added an internal function for checking if a string is a valid Python identifier.
AI: Added a task in Visual Studio Code to export the currently selected Python interpreter path to a file, making it available as “python” and “pip” matching the selected interpreter. This makes it easier to use a specific version with no instructions needed.
AI: Updated the rules to instruct AI to only generate useful comments that add context not present in the code.
Containers: Added template rendering support for Jinja2 (
.j2) container files in our internal Podman tools.Projects: Clarified the current status and rationale of Python 2.6 support in the developer manual.
Debugging: Added experimental flag
--experimental=ignore-extra-micro-passto allow ignoring extra micro pass detection.Visual Code: Added integration scripts for
bashandzshautocompletion of Nuitka CLI options. These are now also integrated into Visual Studio Code terminal profiles and the Debian package.RPM: Included the Python compile script for Linux.
RPM: Removed the requirement for
distutilsin the spec.
Tests
Install only necessary build tools for test cases.
Avoided spurious failures in reference counting tests due to Python internal caching differences. (Fixed in 4.0.3 already.)
Fix, the parsing of the compilation report for reflected tests was incorrect.
Python 3.14: Ignored a syntax error message change.
Python 3.14: Added test execution support options to the main test runner to use this version as well.
Fix, the runner binary path was mishandled for the third pass of reflected compilations.
Removed the usage of obsolete plugins in reflected compilation tests.
Debugging: Prevented boolean testing of
namedtuplesto avoid unexpected bugs.Added the
Testsuffix to syntax test files and disabled “python” mode and spell checking for them to resolve issues reported in IDEs.Fix, newline handling in diff outputs from the output comparison tool was incorrect.
Covered
post-import-codefunctionality with a new subpackage test case.Prevented the program test suite from running an unnecessary variant to save execution time.
macOS: Ignored differences from GUI framework error traces in headless runs in output comparisons.
Reflected test for Nuitka, where it compiles itself and compares its operation has been restored to functional state.
Used the new method to clear internal caches if available for reference counts.
Disabled running nested loops test with Python 2.6.
Containers: Detected Python 2 defaulting containers in Podman tooling.
Cleanups
UI: Fix, there was a double space in the Windows Runtime DLLs inclusion message. (Fixed in 4.0.1 already.)
Onefile: Separated files and defines for extra includes for onefile boot and Python build.
Scons: Provided nicer errors in case of “unset” variables being used, so we can tell it.
Refactored the process execution results to correctly utilize our
namedtuplesvariant, that makes it easier to understand what code does with the results.Quality: Enabled automatic conversion of em-dashes and en-dashes in code comments to the autoformat tool. AI won’t stop producing them and they can cause
SyntaxErrorfor older Python versions, nor is unnecessarily using UTF-8 welcome.Ensured that cloned outline nodes are assigned their correct names immediately upon creation, that avoids inconsistencies during their creation.
Quality: Updated to the latest versions of
blackand adopted a fasterisortexecution by caching results.Quality: Modified the PyLint wrapper to exit gracefully instead of raising an error when no matching files require checking.
Quality: Avoided checking YAML package configuration files twice, since autoformat already handles them.
Quality: Ensured that YAML package configuration checks output the original filename instead of the temporary one when a failure occurs.
Quality: Prevented pushing of tags from triggering git pre-push quality checks.
Quality: Silenced the output of
optipngandjpegoptimduring image optimization auto-formatting.Visual Code: Added the generated Python alias path file to the ignore list.
Quality: Enabled auto-formatting for the Nuitka devcontainer configuration file.
Watch: Avoided absolute paths in compilation to make reports more comparable across machines.
Quality: Changed
mdformatchecks to run only once and silently.Scons: Disabled format security errors in debug mode and moved Python-related warning disables into common build setup code.
Quality: Updated to the latest
deepdiffversion.Scons: Avoided MSVC telemetry since it can produce outputs that break CI.
Debugging: Enhanced non-deployment handler for importing excluded modules.
Split import module finding functionality into more pieces for enhanced readability.
Debugging: Added more assertions for constants loading and checking.
macOS: Dropped the
universaltarget arch.Debugging: Added more traces for deep hash verification.
Summary
This release builds on the scalability improvements established in 4.0, with enhanced Python 3.14 support, expanded package compatibility, and significant optimization work.
The --project option seems usable now.
Python 3.14 support remains experimental, but only barely made the cut, and probably will get there in hotfixes. Some of the corrections came in so late before the release, that it was just not possible to feel good about declaring it fully supported just yet.
Will Kahn-Greene
Bleach 6.4.0 releases -- final release
What is it?
Bleach is a Python library for sanitizing and linkifying text from untrusted sources for safe usage in HTML.
Bleach v6.4.0 released!
Bleach 6.4.0 includes two security fixes, a fix to tinycss2 dependency requirements, and some other things.
See the changes here:
https://bleach.readthedocs.io/en/latest/changes.html#version-6-4-0-june-5th-2026
Bleach v6.4.0 is the final release
I haven't used Bleach on a project in years, but I still had some time to maintain it. That changed about a year ago when I got re-orged into a new role and I haven't had time to do any Bleach work since then.
To recap, Bleach sits on top of html5lib which hasn't been actively maintained in years. It is dangerous to maintain Bleach in that context.
We vendored html5lib so we could make adjustments to the library to keep Bleach going. This is not a sustainable approach, but it was ok for the short term.
Over the years, we've talked about other options:
find another library to switch to
take over html5lib development
fork html5lib and vendor and maintain our fork
write a new HTML parser
etc
None of those are feasible for me.
Bleach has been a solo-maintained project for a while now. The world is crazy and it's much harder to build a team of trusted maintainers now than it was (or at least, it sure feels that way). I don't see any possibility of increasing the maintenance team or passing it to someone else responsibly.
Switching contexts from my regular work to Bleach is really hard. Bleach is complicated, the problem domain is complicated, and there's a lot of nuanced context. I can't just switch gears, spend 15 minutes on Bleach to do something, and then switch back to the rest of my day. I periodically get nag messages about this which are entirely valid, but there's nothing I can do about it. It doesn't feel great.
Then in 2025, Emil, a long-time Bleach contributor, built justhtml which gives us an easy migration path off of Bleach. He even took the time to write a migration guide.
Thoughts and statistics
In 2019, when I stepped down the first time, I wrote a post on stepping down.
In 2023, when I deprecated the project, I wrote a post on Bleach 6.0.0 and deprecation.
From the first commit on 2010-02-18 to today's final commit on 2026-06-05, the Bleach project lasted 16 years, 3 months — 5,951 days, or about 16.29 years.
There were 64 releases.
There were roughly 960 commits.
From 80 roughly contributors
Top 3:
Will Kahn-Greene: 462
James Socol: 182
Greg Guthe: 133
Roughly 5,040 lines of Python code excluding the vendored html5lib.
I was maintainer from October 2015 to now--that's a little under 11 years.
It feels weird to end a project that's outlived many of the Mozilla sites and Python web frameworks it was designed to protect.
What happens now?
This is the end of the project.

Bleach. Last release.
If you're still using Bleach, I think you have three options:
End your project. Maybe you don't need to be maintaining your thing anymore? Use Bleach as your reason to exit and do something different with your time on Earth.
Switch to the sanitizer API. Rework your project to use the sanitizer API.
Swap Bleach out for justhtml. Emil provided a migration guide for switching from Bleach to justhtml.
Good luck with whatever option you choose!
Thanks!
Many thanks to James who created Bleach and gave it a set of first principles that guided our choices for 16 years.
Many thanks to Greg who I worked with on Bleach for a long while and maintained Bleach for several years. Working with Greg was always easy and his reviews were thoughtful and spot-on.
Many thanks to Emil who was a contributor to Bleach for a long while and created justhtml providing Bleach users a migration path.
Many thanks to Jonathan who, over the years, provided a lot of insight into how best to solve some of Bleach's more squirrely problems.
Many thanks to Sam who was an indispensible resource on HTML parsing and sanitizing text in the context of HTML.
Many thanks to all the users and contributors of Bleach!
Where to go for more
For more specifics on this release, see here: https://bleach.readthedocs.io/en/latest/changes.html#version-6-4-0-june-5th-2026
Documentation and quickstart here: https://bleach.readthedocs.io/en/latest/
Source code and issue tracker here: https://github.com/mozilla/bleach/
Real Python
The Real Python Podcast – Episode #298: Reducing the Size of Python Docker Containers
How can you easily reduce the size of a Python Docker container? What are the exceptions you should catch in your code? Christopher Trudeau is back on the show this week with another batch of PyCoder's Weekly articles and projects.
[ Improve Your Python With 🐍 Python Tricks 💌 – Get a short & sweet Python Trick delivered to your inbox every couple of days. >> Click here to learn more and see examples ]
EuroPython Society
EuroPython Society at PyCon US 2026
This year we were back at PyCon US, and this time in sunny Long Beach, California.
We had a booth again, which has quickly become one of our favourite parts of the trip. It&aposs such a great chance to meet folks from other Python communities, catch up with old friends, and put faces to names we&aposve only seen online. People stopped by to chat about EuroPython, pick up stickers, ask about our grants programme, and share what their own local communities are up to. We loved every minute of it.


We also filmed some shorts at the booth, which will be up on our YouTube channel soon! Keep an eye out, there are some lovely conversations in there.
Since EuroPython is celebrating its 25th anniversary this year, we took the chance to talk to community members who have been to many, many EuroPythons over the years. Hearing their stories, the editions they remember most, the friendships that started at one of our conferences, was genuinely moving. It&aposs a good reminder of how much history this community carries with it, and how much of it has been built by people simply showing up year after year.
PyCon US was also where some wonderful people from our community received well-deserved recognition. A huge congratulations to Maria Jose Montreas-Colina, who received an Outstanding PyLady Award for her work with PyLadies and the wider community. Maria is part of our team and helps look after PyLadies and community matters at EuroPython. Congratulations also to Rodrigo Girão Serrão for receiving the Community Service Award for his contributions to the community. Rodrigo works on our programme and sprints.
Thank you both for everything you do. 💛


Long Beach itself was a lovely city. Palm trees, warm weather, the ocean nearby. A very different vibe from the usual conference cities, and a really lovely backdrop for a week of Python.
A big thank you to the PyCon US organisers for having us, and for making space for the wider Python world to come together. And a thank you to everyone who stopped by the booth to say hello, it was a pleasure meeting you.
See you next year, and we hope to see many of you in Kraków for EuroPython 2026!


Bob Belderbos
How to Update Multiple Page Elements from One htmx Request
A button submits code, tests run, feedback appears. Standard htmx. But the submissions dropdown stays stale; the new submission is in the database, just not in the dropdown. One request, two elements to update.
The problem: one request, two things to update
On our Rust platform, each exercise page has a "Run Tests" button. It posts the editor code to a Django view, which compiles and runs the tests, then swaps a pass/fail panel into a #feedback div.
Next to the editor there is a dropdown of your past submissions. Run the tests, and a new submission gets saved server-side. But the dropdown stayed stale until you reloaded the page. The new submission was there in the database, just not in the <select>.
So now I have one request that needs to update two unrelated parts of the page: the feedback panel (the htmx target) and the submissions dropdown (somewhere else entirely in the DOM).

First instinct: write JavaScript to read the response, build a new <option>, prepend it to the select. That works until you remember the dropdown also enforces a max number of submissions, drops duplicates, and orders newest-first. Replicate that logic in the browser and you now have two sources of truth that drift apart the first time you change the server rule.
Htmx out-of-band swaps
htmx has a feature for exactly this: out-of-band swaps.
The hx-swap-oob attribute allows you to specify that some content in a response should be swapped into the DOM somewhere other than the target, that is "Out of Band". This allows you to piggyback updates to other element updates on a response.
htmx swaps any element carrying hx-swap-oob into its matching target on the page, separately from the main swap. One response, many updates.
First I pulled the submissions dropdown options into a partial so the page and the view render them identically:
<!-- _submission_options.html -->
<option disabled selected>Submissions / Reset</option>
{% for submission in submissions %}
<option value="{{ submission.unique_hash }}">{{ submission.created_at|date:"Y-m-d H:i" }} {% if submission.ok %}(OK){% else %}(Failed){% endif %}</option>
{% endfor %}
<option value="reset">Reset</option>The page includes it inside the <select id="submissions">. The view renders the same partial and tags it for an out-of-band swap:
from django.template.loader import render_to_string
from django.utils.html import escape
def validate(request):
# ... run the tests, save the submission ...
submissions = Submission.objects.filter(
exercise=exercise, user=user
).order_by("-created_at")
options = render_to_string(
"_submission_options.html", {"submissions": submissions}
)
return HttpResponse(
f"""
<div class="...">{message}</div>
<pre>{escape(output)}</pre>
<div hx-swap-oob="innerHTML:#submissions">{options}</div>
"""
)The first part of the response swaps into #feedback as usual. htmx spots the hx-swap-oob element, pulls it out, and applies it to #submissions instead. The button HTML only knows about #feedback. The view decides what else to update. Whoever owns the data controls how it renders.
A second example: progress bars that update themselves
On the Python platform the same trick drives the learning-path progress widget. Passing an exercise recomputes your progress along every path it belongs to and swaps the bars into a sidebar, from the same request that renders the pass/fail panel:
if ok:
paths_html = ""
for path in bite.bite_paths.prefetch_related("bites"):
# ... compute completed / total / pct for this path ...
paths_html += render_progress_bar(path, completed, total, pct)
extra_html += (
f'<div id="learning-paths-progress" hx-swap-oob="innerHTML">{paths_html}</div>'
)The two examples aim at their targets differently. The progress widget uses a bare hx-swap-oob="innerHTML": htmx swaps the fragment into whatever element already shares its id. The dropdown uses the selector form, hx-swap-oob="innerHTML:#submissions", so the carrier <div> can target the <select id="submissions"> without needing to share its id.
Use innerHTML instead of the default hx-swap-oob="true". true replaces the whole element (its outerHTML), which for the <select> throws away the htmx listener attached to it. innerHTML keeps the element and swaps only its children, so the listener survives and the options refresh underneath it.
Here's the progress bars before and after passing an exercise. The bars update via out-of-band swap from the same response that renders the pass/fail feedback:


You might spot the submissions dropdown in these shots and wonder why it is not OOB-swapped here too, like on the Rust platform above. It is the boundary of the technique: OOB swap is solid when you replace an element's whole inner content (innerHTML:#submissions), but inserting a single new <option> means wrapping it in a <template> so the browser does not strip it, and htmx 2.x handled that case unreliably. So here the one-option insert runs through a small htmx:afterSwap listener instead. The view writes the new submission into a hidden div, and the listener reads it and prepends the <option>:
document.addEventListener("htmx:afterSwap", () => {
const newSub = document.getElementById("new-submission");
if (!newSub) return;
const select = document.getElementById("submissions");
const opt = document.createElement("option");
opt.value = newSub.dataset.hash;
opt.textContent = newSub.dataset.label;
select.insertBefore(opt, select.children[1]); // after the placeholder
select.value = opt.value;
});One honest cost: this adds a query. After saving, the view re-fetches the submissions to render the partial. The alternative is re-implementing state changes in pure JavaScript, creating behavior in two places. With out-of-band swaps you drive the logic from the view, all in one place. One more query, but less code and a more maintainable solution.
Whoever fetches the data should render it. Keep the query and the template together.
For more on hypermedia-driven applications, see this great book: Hypermedia Systems.
Seth Michael Larson
Is the Super Smash Bros. Brawl donut from Mister Donut?
Happy Donut Day (and #FediDonutFriday) to those who celebrate! 🍩 Present and Correct shared a link to the Mister Donut museum on Bluesky and upon clicking through I was greeted with a familiar face: a chocolate ring donut.
Strangely, I've seen this chocolate ring donut before: from the hours staring at sprite-sheets from the Super Smash Bros. and Kirby Air Riders franchises. That donut looked just like the one from Super Smash Bros. Brawl.
“But Seth”, I hear you say, “chocolate ring donuts all look the same anyway!” Maybe... and yet...


Funnily enough, the Render96 wiki, which collects origins for artwork such as Super Smash Bros. Melee and Brawl, lists the donut as one of the few foods where the origin is not known. Could this donut also be a Mister Donut? We'll probably never know!
Thanks for keeping RSS alive! ♥ What to do next? Share your thoughts with me on Mastodon, Bluesky, or email. I try to reply to everyone!Browse the blog archive. Check out my blogroll. Or maybe go outside (best option)?
June 04, 2026
The Python Coding Stack
Down The Iterator Rabbit Hole
You know that street game where the performer (con artist?) has three opaque cups and a small ball. He places the cups upside down on the table, with the ball under one of the cups. He quickly shuffles the cups around and then asks the player to guess which cup has the ball. You’ve seen the game on TV, even if you’ve not seen it in real life.
Following what’s happening when you have a chain of iterators in Python can feel like playing that game. But, unlike the street game, there are no scams when you’re playing the iterator game. Let’s make sure you’ll always win.
I’ll keep this article short. I wrote many articles about iterables and iterators. If you need to refresh your memory, have a look at The Anatomy of a for Loop and A One-Way Stream of Data • Iterators in Python (Data Structure Categories #6).
Follow The Data in a Chain of Iterators
Let’s keep the example simple. Start with this list in a REPL session:
A list is iterable. You can create an iterator from any iterable. Let’s create an iterator from this list:
The built-in function iter() creates an iterator from an iterable. Iterators don’t contain data. They don’t create copies of the data. They’re lightweight objects that create a stream. They’ll fetch data from the original source, which is the list boring_numbers in this case, as and when needed.
Iterators can only fetch an item once. So, they’re a one-way stream. Once you use an item, it’s gone from the iterator – but not from the original list, which remains unchanged.
Therefore, first_iter is an iterator that relies on data from the list boring_numbers. But let’s not fetch any items from the first_iter iterator. Not yet, anyway.
Create a second iterator. This time, you’ll use a generator expression. Generators are iterators, so you create a second iterator with this code:
Note that the expression on the right-hand side of the equals sign is enclosed in parentheses – the round ones, to be clear. This is a generator expression, which creates a generator iterator. Read Pay As You Go • Generate Data Using Generators (Data Structure Categories #7) for more on generators.
As we said, generators are iterators.
The second_iter iterator generates data from first_iter, which is itself an iterator. Iterators are also iterable, which is why you can use them directly in a for clause or anywhere else you’d generally use an iterable. The second_iter iterator will yield the values as floats. But you’ve not yielded any value from this iterator either. Not yet.
Let’s go a step further and create a third iterator, which is also a generator in this case. You build this third iterator from the second one, second_iter:
The generator iterator third_iter yields the sum of 0.5 and the value yielded by second_iter.
Incidentally, I used a “standard” iterator and two generator iterators in this example. However, for the journey we’re following in this article, it doesn’t matter whether we’re using a basic iterator or a generator iterator. If you prefer, you can repeat this exercise with iterators you get from iter() directly.
Don’t Blink • Follow the Data
You started with a list called boring_numbers. This data structure contains* the data. It’s where the data lives. We’ll be following the data in this section. So it’s important to know where it’s stored!
*Note: Lists, like all data structures, don’t really contain data in the purest sense of the word. See What’s In A List—Yes, But What’s Really In A Python List for more on this. But in general, it’s fine to talk about a list ‘containing’ items of data.
You then create three iterators. The first uses data from boring_numbers. The second iterator uses data from the first. And the third iterator uses data from the second.
But you haven’t tried to fetch any value from any of the iterators yet.
Let’s look at what each iterator is doing at the moment before you fetch any values. The first iterator, first_iter, is pointing at the first item in boring_numbers. It’s ready to read this value and yield it.
The second iterator, second_iter, is pointing at the first item in first_iter. But first_iter doesn’t have any data. Iterators don’t have their own data. But that’s OK. Whenever second_iter needs to fetch the value, it will ask first_iter to fetch and yield its “first” value. I put “first” in quotation marks because you’ll see later that this may or may not be the first value.
Finally, third_iter is pointing at the first item in second_iter. The same logic applies. When third_iter needs the first item, it will ask second_iter for its “first” item, and second_iter will need to ask first_iter for its “first” item. And first_iter is pointing at the first item in the list boring_numbers.
Are you with me? Let’s complicate things a bit…
Note how your code so far includes the following lines:
None of the iterators has yielded any value. For now.
Let’s jumble things up and start by fetching the first value from second_iter:
You ask for the next value in second_iter, which is the first one since you haven’t yielded any values yet.
As you’ve seen earlier, second_iter needs the first value from first_iter. So, behind the scenes, Python calls next(first_iter), which yields the first item from boring_numbers.
So, first_iter reads the first value from boring_numbers, which is the integer 1, and it yields it to second_iter, which then yields the transformed version to the REPL as the return value of next(second_iter). That’s why the output is the float 1.0. The first iterator, first_iter, now moves to point at the second item in boring_numbers, ready for when it’s needed.
Note that boring_numbers doesn’t change in this process. The first item in boring_numbers remains there. It doesn’t disappear.
So far, so good?
Continue in the same REPL session and try the following:
You ask third_iter to give you its “next” value. You haven’t used third_iter anywhere so far. So, you might expect it to yield the “first” value.
And it does.
But its interpretation of what’s the “first” item may be different to what you expect.
Let’s follow the data. When you call next(third_iter), the third iterator asks second_iter for its next item. The second iterator, second_iter, relies on first_iter, so it asks first_iter for its next item. And first_iter, as you may recall, is currently pointing at the second item in boring_numbers, which is the integer 2.
So:
The first iterator
first_itergets the integer2fromboring_numbersand yields it tosecond_iter. Andfirst_iternow points at the third item inboring_numbers.Then,
second_itertransforms this value into a float and yields2.0tothird_iter.Finally,
third_iteradds0.5to this value and yields2.5, which is what you see displayed in the REPL.
When you called next(second_iter) earlier in the code, you used up the first item in second_iter, which in turn used up the first item in first_iter. Since this first value is gone and since third_iter depends on the data yielded by second_iter and first_iter, the earlier call to next(second_iter) also affected the iterator that’s downstream, third_iter.
What will happen if you call next(first_iter) now? Try to follow the data in your head before trying it out or reading on.
.
.
Have you worked it out?
.
.
Let’s run the code:
Although it’s the first time you explicitly use first_iter in your code, you already used two of its values when your code yielded values from iterators downstream. Therefore, the next item in first_iter is the third item in boring_numbers, the integer 3.
Let’s finish with one more expression, still running in the same REPL session:
You call next(third_iter), which asks second_iter for its next item. And second_iter asks first_iter for its next item. At this stage in the process, first_iter is pointing at the fourth item in the original source of data, which is the list boring_numbers. That’s why the output is 4.5.
Independent Iterators
Consider the following code, which is similar to the one you wrote above but has one extra line:
The iterators first_iter and another_first_iter both use the same source of data, boring_numbers. However, they are independent iterators. Note that when you use up some of the elements in first_iter, the independent another_first_iter is not affected. The first time you ask for the first item in another_first_iter, you get the integer 1.
Final Words
Iterators don’t contain data. They rely on data that’s stored elsewhere. But you can have a chain of iterators, each asking the previous one to yield a value. Weird things can happen if you’re not careful. But now you know how to follow the data when you have a chain of iterators.
As a rule of thumb, if you create an iterator that depends on another iterator, you should only use the final iterator to avoid these issues. So, in the example above, you should only yield values from third_iter.
Have a play with this example and make your own chains of iterators, too. And once you’re comfortable with this, get ready to be confused again with my next article, which will discuss itertools.tee()!
And next time you pass by someone in the street offering to let you play the three-cups-and-ball game, don’t feel overconfident because of your iterator knowledge – it won’t help you find the ball.
Code in this article uses Python 3.14
The code images used in this article are created using Snappify. [Affiliate link]
Join The Club, the exclusive area for paid subscribers for more Python posts, videos, a members’ forum, and more.
For more Python resources, you can also visit Real Python—you may even stumble on one of my own articles or courses there!
Also, are you interested in technical writing? You’d like to make your own writing more narrative, more engaging, more memorable? Have a look at Breaking the Rules.
And you can find out more about me at stephengruppetta.com
Further reading related to this article’s topic:
A One-Way Stream of Data • Iterators in Python (Data Structure Categories #6)
Pay As You Go • Generate Data Using Generators (Data Structure Categories #7)
Appendix: Code Blocks
Code Block #1
boring_numbers = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10]Code Block #2
# ...
first_iter = iter(boring_numbers)Code Block #3
# ...
second_iter = (float(number) for number in first_iter)Code Block #4
# ...
third_iter = (num + 0.5 for num in second_iter)Code Block #5
boring_numbers = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10]
first_iter = iter(boring_numbers)
second_iter = (float(number) for number in first_iter)
third_iter = (num + 0.5 for num in second_iter)Code Block #6
# ...
next(second_iter)
# 1.0Code Block #7
# ...
next(third_iter)
# 2.5Code Block #8
# ...
next(first_iter)
# 3Code Block #9
# ...
next(third_iter)
# 4.5Code Block #10
boring_numbers = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10]
first_iter = iter(boring_numbers)
another_first_iter = iter(boring_numbers)
second_iter = (float(number) for number in first_iter)
third_iter = (num + 0.5 for num in second_iter)
next(second_iter)
# 1.0
next(third_iter)
# 2.5
next(first_iter)
# 3
next(third_iter)
# 4.5
next(another_first_iter)
# 1For more Python resources, you can also visit Real Python—you may even stumble on one of my own articles or courses there!
Also, are you interested in technical writing? You’d like to make your own writing more narrative, more engaging, more memorable? Have a look at Breaking the Rules.
And you can find out more about me at stephengruppetta.com
Real Python
Quiz: How to Read User Input From the Keyboard in Python
In this quiz, you’ll test your understanding of How to Read User Input From the Keyboard in Python.
By working through this quiz, you’ll revisit the input() function, type conversion, error handling with try and except, the getpass module for hidden input, and the PyInputPlus library for automatic validation.
[ Improve Your Python With 🐍 Python Tricks 💌 – Get a short & sweet Python Trick delivered to your inbox every couple of days. >> Click here to learn more and see examples ]
Python Software Foundation
PSF Strategic Plan 2026 Draft: Open for Community Feedback
In May, we shared the high-level goals of the Python Software Foundation's (PSF) strategic plan and asked for your commentary. Today we are publishing the full draft and opening a three-week community feedback window.
We welcome you to review the full PSF Strategic Plan Community Draft 2026 document, also embedded below.
The feedback window closes on June 25, 2026, End Of Day, Anywhere on Earth. The PSF Board will carefully review all input, use it to refine the final version of the strategic plan, and aims to hold a vote to adopt it in a future board meeting.
What's in the full draft
The earlier blog post covered the six organizational goals and four program goals at a high level. The full draft goes deeper: each program goal includes specific strategic objectives, and the organizational goals include tactical ideas the board developed during the planning process. These tactical ideas are starting points for strategic discussion, not commitments.
This is the first post in a short series. Individual board members will share posts that go into specific parts of the plan in more depth. We want the plan to speak for itself, so these posts will draw directly from the document rather than rewriting it.
What we heard at PyCon US
At PyCon US 2026, the PSF Board held its on-site board meeting, with a portion of that time dedicated to strategy. We also discussed the strategic plan at the Members Lunch, a dedicated Open Space session, and in conversations throughout the conference.
The topic of financial sustainability came up repeatedly, and we hear you. The community is waiting for updated financial information, and typically the Members Lunch at PyCon US is where those details are shared. Staffing changes in our accounting functions made that impossible this year. Publishing the full picture is a priority, and we will share an update as soon as we can. The high-level view is that the PSF is stable for now, but we cannot continue on the current path without making meaningful changes. The strategic plan and the PSF's financial outlook are connected, and we understand that context matters. We are committed to being transparent about both.
We also noticed that conversations naturally moved toward implementation ("How will you do this?"). For this feedback round, we are asking you to focus on the direction itself. Are these the right goals? Are the objectives the right ones? Is anything important missing? Implementation will be shaped by PSF staff over time, and there will be opportunities to weigh in on that, too.
How to give feedback
- Email strategy@python.org to share detailed or private feedback. This is the best way to reach us.
- Discuss thread for open conversation.
- PSF Board Office Hours on the PSF Discord on:
The feedback window closes on June 25th. After that, the board will review all feedback received and decide what changes to make to the strategy document in response.
Thank you for your time. We’re working on this strategic plan because the Python community deserves a PSF that's deliberate about where it's headed. Your input makes that possible, and we’re grateful for your help.
Jannis Leidel, PSF Board Chair, on behalf of the PSF Board of Directors
Adrarsh Divakaran
Building AI Agents in Python
2026 is shaping up to be a big year for AI agents. We are seeing more products where the AI not only answers a question but also does some work for the user.
You have probably used ChatGPT or a similar AI tool to answer a question, help with writing, or explain some code. You type something, the AI responds, and the conversation goes back and forth. That is powerful, but it is also limited. The AI is essentially stuck in a chat box - it can only talk to you; it cannot do anything on your behalf.
AI agents change that. An agent is an AI that can actually take actions - browse the web, read and write files, run code, call APIs, and more. It does not just answer your question; it works toward a goal, step by step, using whatever tools it needs. Tools like Lovable, Cursor, and Claude Code are examples of this in practice.
In this article, we will explore the concepts behind building an AI agent in Python. We will use the OpenAI Python SDK (Responses API) for the examples, but the same ideas can be generalized to any other LLM SDK. We will use a low-level SDK with minimal abstractions so we can observe and implement most of the agent’s behavior on our end.
TL;DR
This tutorial explains how AI agents work by building a simple one in Python.
We will cover the core pieces: LLMs, prompts, context, memory, the agent loop, tools, MCP, and skills:
| Component | What it does |
|---|---|
| LLM | Acts as the reasoning engine that understands the user request and decides what to do next. |
| System prompt | Defines the agent’s role, behavior, boundaries, and response style. |
| Context window | Controls how much information the model can see at once, including prompts, history, tool results, and files. |
| Memory | Helps the agent remember useful information across steps or conversations. |
| Agent loop | Repeats the process of thinking, acting, observing results, and deciding the next step. |
| Tool calling | Lets the agent use external functions such as APIs, web search, file access, or code execution. |
| MCP | Provides a standard way to connect agents to reusable tools and data sources. |
| Skills | Package reusable instructions, workflows, examples, and scripts for specific tasks. |
What are Agents?
An AI agent is an AI system that can autonomously plan and execute multi-step actions toward a goal.
To understand agents, it helps to first understand what is powering them under the hood - a large language model, or LLM. For example, ChatGPT is a product built on top of OpenAI GPT LLMs. When you type a message and get a response, an LLM is doing the heavy lifting. It takes text as input and generates text as output.
On their own, LLMs are impressive but limited. They can only respond with text. They cannot open your browser, read a file on your computer, or send an email. They also do not know what happened yesterday, because their knowledge comes from training data with a cutoff date, not a live connection to the world.
Agents fix this by giving LLMs access to tools. A tool is just a function your code exposes to the model - something like “search the web” or “read this file.” The model can decide to call a tool when it needs to, and your code actually runs it. This turns a passive text generator into something that can act.
A good way to see the difference is to compare using ChatGPT with using Claude Code for a coding task. With ChatGPT, you describe the problem, copy the suggested code, paste it into your editor, run it, copy the error back, and repeat. The model has no idea what is actually in your project. Claude Code is different - it is powered by an LLM but also has access to tools like bash and file reading. You describe what you want, and it reads your files, writes code, runs tests, and fixes errors on its own. You just watch and steer.
The simplest way to understand an agent is:
- The user gives a goal.
- The model decides what step to take.
- The agent runs that step using a tool.
- The model looks at the result.
- The process continues until the task is complete.
This is different from a normal chatbot. A chatbot mainly responds. An agent can respond and act.
In a simple agent, the model may only call one tool and return the result. In a more capable agent, the model may make a plan, call multiple tools, observe the results, adjust the plan, and continue until the task is complete.
Before we build this kind of system, we need to choose the model that will drive it.
LLMs
LLMs are trained on massive amounts of text data - entire open source repositories on GitHub, books, articles, websites, and more. Through training, the model learns patterns in language well enough to generate coherent, useful responses. The scale of this training is what makes them surprisingly capable across such a wide range of tasks.
At their core, LLMs are text-in, text-out systems. You send them a block of text (called a prompt), and they generate a response. Everything that happens - reasoning, answering questions, writing code, making decisions - is expressed through that text interface. When an agent calls a tool, it is really the model writing out a structured text request, and your code intercepts that and actually runs the function.
The key limitation to keep in mind: LLMs only know what they were trained on. They have no awareness of events after their training cutoff and no way to look things up in real time unless they are given a tool to do so. This is part of what makes tools so valuable - they extend the model’s reach into the real world.
Choosing an LLM
For an AI agent, the LLM is its brain. The quality of the model affects how well the agent understands instructions, chooses tools, handles errors, and completes multi-step tasks.
At the same time, the most powerful model is not always the right choice. We also need to think about cost, speed, context window, reasoning ability, and where the model is hosted.
Benchmarks
Benchmarks are standardized tests used to compare the performance of different models. For coding tasks, there is SWE-bench. For general reasoning, there is MMLU. Each benchmark tests the model on a specific type of problem and gives it a score. A higher score generally means the model will perform better on that type of task.
Benchmarks are a useful starting point when choosing a model, but they are not the whole story. A model that scores well on a benchmark may still behave unexpectedly in your specific use case, so it is always worth testing with your actual workload.
Costs
Choosing the best-scoring model from a benchmark may not always be the most intelligent decision.
Cost is a real factor, especially at scale. Most providers charge per token, which is the basic unit of text the model processes. A token is roughly four characters, or about three-quarters of a word on average. Both what you send to the model (input) and what it generates back (output) count toward your token usage.
For an agent that runs multiple steps in a loop, token usage adds up quickly. A good approach is to start with a capable model and then see if a smaller or cheaper one can do the same job well enough. Sometimes a smaller model handles simple tasks just fine.
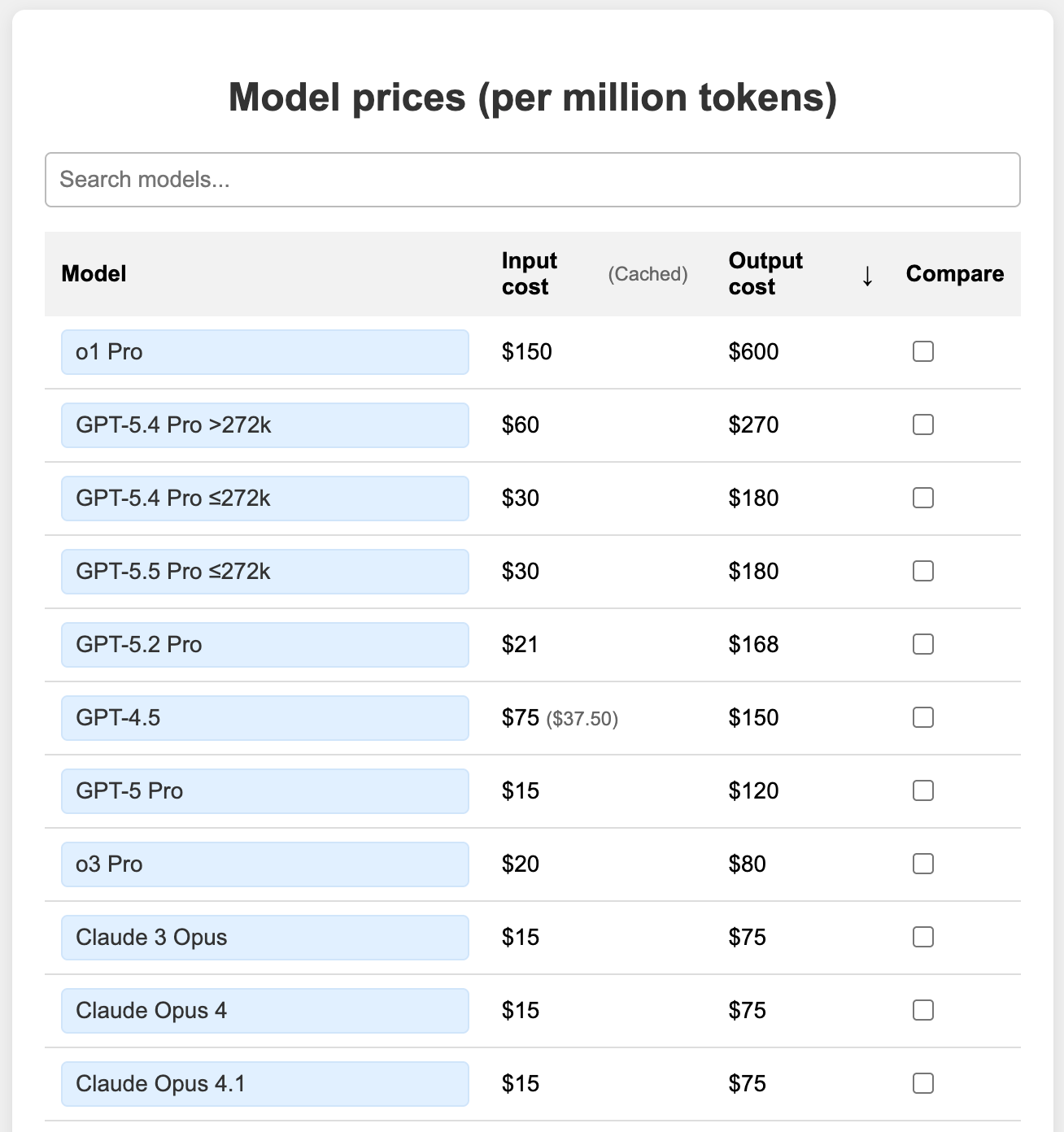 (Model costs table from https://github.com/simonw/llm-prices)
(Model costs table from https://github.com/simonw/llm-prices)
Reasoning Level
Some models are designed to think before they answer. These reasoning models break complex problems into smaller steps internally, often called reasoning traces. You can think of it as the model working through a scratchpad before writing its final response. This can improve performance for tasks that need planning, debugging, tool use, or careful decision-making.
More reasoning effort usually means higher cost, higher response time, and better accuracy for complex tasks.
Not every request needs high reasoning. If the task is simple, we can use a lower reasoning level or a cheaper model. If the task involves multiple steps, unknown errors, or important decisions, more reasoning can be useful.
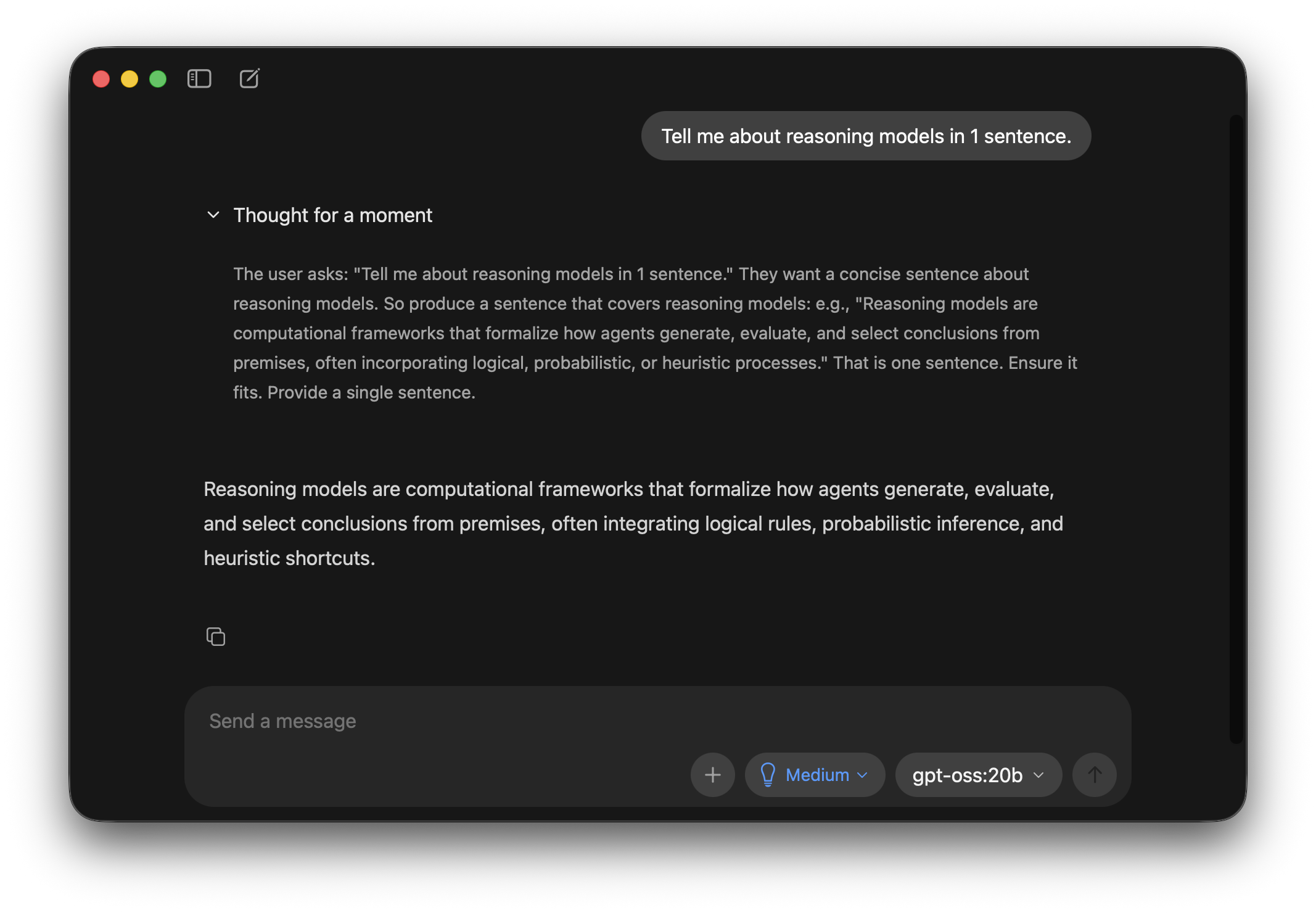 (Conversation with GPT-OSS LLM showing reasoning/thought traces)
(Conversation with GPT-OSS LLM showing reasoning/thought traces)
Hosted vs Local
Most people start with a hosted model - one that runs on a provider’s servers and is accessed via an API. These are easy to set up, well-maintained, and generally the most capable options available. The trade-off is that you pay per token, and your data is processed by a third party.
There are also open models that can run entirely on your own machine/server. They can avoid per-token API costs and give you more control over data. The downside is that they require capable hardware and are generally less powerful than the best hosted models today. That said, local models are getting better quickly. Previous generation frontier capabilities are being replicated in the next generation of local models, and this gap will continue to close. Examples of open-weight models that can be self-hosted, depending on hardware and quantization, include Gemma 4 series and Kimi K2.6.
There are already decent local coding models that people use for simple code generation and verification. In the coming years, this will improve, and stronger models will become available on consumer devices.
Hosted models are still easier to use for many applications. They usually provide better quality, higher reliability, larger context windows, and managed infrastructure.
Local models give more control over data, cost, and deployment. But they also require more setup, hardware, monitoring, and optimization.
Configuring the LLM
Once you have picked a model, there are two things you set up before the agent starts running: the system prompt and the context window.
System Prompt
A system prompt is the model’s top-level instruction that guides its behavior during a conversation.
It can set rules such as:
- what role the AI should play
- what tone it should use
- what it should or should not do
- how it should handle tools
- how it should handle safety and user requests
- how it should format the final answer
For an agent, the system prompt is very important. It tells the model how to behave while using tools. It can also define boundaries, such as asking for permission before destructive actions or avoiding actions outside the user’s request.
Let’s see an example of this in practice:
import os
if __name__ == '__main__':
from openai import OpenAI
client = OpenAI(api_key=os.environ["OPENAI_API_KEY"])
response = client.responses.create(
model="gpt-5.4-mini",
input=[
{
"role": "system",
"content": "You are a friendly Python tutor. Refuse all requests unrelated to Python coding",
},
{
"role": "user",
"content": input("Enter your Python question: "),
},
],
)
print(response.output_text)
In the above script, we initialize an OpenAI client and use client.responses.create to send a message to gpt-5.4-mini model. The system prompt is specified in the input list as the first entry. "role": "system" designates the entry as the system prompt. In the above example, the model is instructed to act as a Python tutor and refuse requests unrelated to Python. As the next entry, we accept the user prompt via input() and pass it to the LLM for answering.
If the script is run and any unrelated queries are passed to the LLM, we get a refusal response similar to the below one:
Enter your Python question: How many states are there in the US?
Model response: I’m here to help with Python coding questions only. If you have a Python-related question, feel free to ask!
Even though the underlying large language model knows the answer to the user’s query, it refuses to answer as per direction in the system prompt.
Context Window
The context window is the model’s working memory. It is the amount of information the model can see in one request.
The context can include the user message, conversation history, system prompt, tool results, files, documentation, and any other information we provide.
Most of the latest flagship models support up to 1M tokens, which is roughly 750,000 words or about 15 books. Older models like GPT-4 series models had a 128K token window, around 2 books’ worth. For agents that run long tasks or work with large documents, context window size matters a lot. When the context fills up, older information gets dropped, which can cause the agent to lose track of earlier steps in a long task.
A larger context window is useful, but it is not free. More context usually means more cost and slower responses. Also, just because a model can accept a lot of context does not mean every token is equally important.
Good agents manage context carefully. They include what is needed, summarize old information, and avoid filling the context with unnecessary data.
Once we understand the model and its context window, the next question is what the agent should remember across steps and conversations.
Memory
Memory helps an agent remember useful information.
Short-term memory helps the agent remember what the user said earlier in the same conversation. This usually lives inside the context window.
Let’s consider an example. The snippet below accepts a user query inside a loop and sends it to a model to get the response:
import os
if __name__ == '__main__':
from openai import OpenAI
client = OpenAI(api_key=os.environ["OPENAI_API_KEY"])
while True:
user_query = input("You: ")
if user_query.lower() in ["exit", "quit"]:
break
response = client.responses.create(
model="gpt-5.4-mini",
input=user_query,
)
assistant_reply = response.output_text
print(f"Model: {assistant_reply}")
The code works, but there are issues:
You: Tell me about Taj Mahal in 1 sentence
Model: The Taj Mahal is a magnificent white marble mausoleum in Agra, India, built by Emperor Shah Jahan in memory of his wife Mumtaz Mahal, and is one of the world’s most famous symbols of love.
You: When was it built?
Model: I can help, but I need to know **what “it” refers to**.
Please share the name, photo, or location of the building/structure/object, and I’ll tell you when it was built.
As seen from the transcript, the model fails to answer the user’s follow-up prompt. This is because, we did not implement short term memory. For the model to be able to respond to follow-ups properly, we need to store and pass the conversation history to LLM calls. The snippet improves on the above script with short term memory implementation:
import os
if __name__ == '__main__':
from openai import OpenAI
client = OpenAI(api_key=os.environ["OPENAI_API_KEY"])
conversation_history = []
while True:
user_query = input("You: ")
if user_query.lower() in ["exit", "quit"]:
break
conversation_history.append({
"role": "user",
"content": user_query,
})
response = client.responses.create(
model="gpt-5.4-mini",
input=conversation_history,
)
assistant_reply = response.output_text
print(f"Model: {assistant_reply}")
conversation_history.append({
"role": "assistant",
"content": assistant_reply,
})
We introduced a conversation_history list that stores previous messages. User messages are appended to this list with "role": "user" and model responses are appended with "role": "assistant". This way, whenever a request is sent to the model, it gets the entire message history through the input argument and will be able to respond to follow-up prompts correctly.
You: Tell me about Taj Mahal in 1 sentence
Model: The Taj Mahal is a stunning white marble mausoleum in Agra, India, built by Emperor Shah Jahan in memory of his wife Mumtaz Mahal.
You: When was it built?
Model: It was built between 1632 and 1653.
Long-term memory stores information beyond one conversation and persists even after the current chat or task ends. This is useful when you want the agent to remember user preferences, past decisions, or domain-specific facts across sessions. Common approaches include RAG (retrieval-augmented generation), where relevant information is fetched from a database and added to the context as needed, and built-in memory systems like ChatGPT Memories, where key facts are stored and automatically recalled in future conversations.
Agent Loop
The agent loop is the core flow of an agent.
A simple loop looks like this:
- User sends a message.
- Agent adds the message to the conversation context.
- Agent sends the context and system prompt to the LLM.
- LLM decides what to do next.
- If needed, the LLM calls a tool.
- Agent runs the tool and sends the result back to the LLM.
- LLM decides whether more steps are needed.
- When done, the LLM generates the final response.
- Agent sends the response to the user.
This loop is what makes agents feel different from normal chatbots. A chatbot usually gives one response. An agent can act, observe, and continue.
In practice, the intermediate steps are where the interesting work happens. The model may call a tool, wait for the result, process that result, decide to call another tool, and keep going before it gives a final answer. The loop runs as many times as needed until the model decides the task is complete or the user stops it. This brings us to tools - what they are and how they actually work.
Tool Calling
Tools are external capabilities that the agent can use.
Tools (also called functions) let an AI agent do things beyond generating text. They can be used to take actions or get information.
Examples of tools:
- search the web
- call an API
- read files
- edit files
- run code
- send emails
- query a database
- create calendar events
The agent chooses a tool when needed. The tool has a name, a description, and input parameters. The model decides which tool to call and what arguments to pass.
Tool descriptions are important. If a tool description is unclear, the model may call it at the wrong time or pass the wrong input. We should describe tools in simple language and make their inputs strict.
Here is an important detail: the model does not run the tool itself. When it decides to use a tool, it outputs a structured request with the tool name and the arguments it wants to pass. Your code intercepts this, runs the actual function, and passes the result back to the model. The model then reads the result and decides what to do next. This back-and-forth between the model and your code is what makes the agent loop so powerful.
Let’s see an example of tool calling in action:
import json
import os
from dotenv import load_dotenv
load_dotenv()
def get_weather(location):
return {
"location": location,
"temperature": "24 C",
"condition": "Sunny",
"humidity": "52%",
"wind": "11 km/h",
}
if __name__ == '__main__':
from openai import OpenAI
client = OpenAI(api_key=os.environ["OPENAI_API_KEY"])
tools = [
{
"type": "function",
"name": "get_weather",
"description": "Get the current weather for a destination.",
"parameters": {
"type": "object",
"properties": {
"location": {
"type": "string",
"description": "The city or destination, e.g. Paris or Tokyo",
}
},
"required": ["location"],
"additionalProperties": False,
},
"strict": True,
}
]
input_list = [
{
"role": "system",
"content": "You are Safar, a travel planning AI agent",
},
{
"role": "user",
"content": input("Ask you travel questions: "),
},
]
response = client.responses.create(
model="gpt-5.4-mini",
input=input_list,
tools=tools,
tool_choice="required",
)
print("The model responded with:")
print(response.output)
input_list += response.output
for item in response.output:
if item.type != "function_call":
continue
if item.name == "get_weather":
args = json.loads(item.arguments)
print(f"The model wants to call get_weather with: {args}")
weather = get_weather(args["location"])
print(f"The local Python function returned: {weather}")
input_list.append({
"type": "function_call_output",
"call_id": item.call_id,
"output": json.dumps(weather),
})
print("Sending the tool result back to the model")
final_response = client.responses.create(
model="gpt-5.4-mini",
input=input_list,
tools=tools,
)
print("Final answer:")
print(f"Model response: {final_response.output_text}")
In the tools list, we have defined a function named get_weather according to OpenAI function calling guidelines and have specified the parameters that the model accepts using the parameters key. This definition follows JSON Schema specification.
Since, we add this tools list when making calls to OpenAI, the model will know that it has access to a weather tool and will be able to request a function call when needed.
In the script, you can see that when we receive a response from the model, we always check if the response type is a function call or not (item.type != "function_call") and if the response is a request to call get_weather tool, we call the get_weather() Python function and send it back to the model:
weather = get_weather(args["location"])
input_list.append({
"type": "function_call_output",
"call_id": item.call_id,
"output": json.dumps(weather),
})
Let’s run the script and ask the agent a question that would require a weather tool call:
Ask you travel questions: Sunscreen needed in Goa?
The model responded with:
[ResponseFunctionToolCall(arguments='{"location":"Goa"}', call_id='call_X9OBZhGwT3yhfmTAOclefWE8', name='get_weather', type='function_call', id='fc_05ba95ec38f46f7f006a17ce9e3bb0819a9a0b430001f7bd91', namespace=None, status='completed')]
The model wants to call get_weather with: {'location': 'Goa'}
The local Python function returned: {'location': 'Goa', 'temperature': '24 C', 'condition': 'Sunny', 'humidity': '52%', 'wind': '11 km/h'}
Sending the tool result back to the model
Final answer:
Model response: Yes — sunscreen is a good idea in Goa. It’s sunny there right now, so UV exposure can be strong even if it feels pleasant.
Quick tips:
- Use broad-spectrum SPF 30+ (SPF 50 if you’ll be at the beach a lot)
- Reapply every 2 hours, and after swimming/sweating
- Don’t forget ears, neck, hands, and feet
- A hat and sunglasses help too
If you want, I can also suggest a Goa beach-day packing list.
For our query, the model initially responds with a ResponseFunctionToolCall item. This requests our get_weather function to be called with location argument set as Goa.
Responding to this request, our script executes the function call and sends the function call response back to the model for getting the final response. The function call always returns temperature as 24 degree Celsius with condition as sunny. Trusting this data, the model produces its final response, suggesting the user to use a sunscreen.
The weather function defined in the above script is not a very useful one, it returns a hardcoded weather data for all requests. In a practical scenario, the function should make an actual call to a real Weather API to fetch data.
The above script illustrates the concept of an agent loop. Even though the example involves just one user request and model response, the agent takes intermediary steps (tool calls) before returning the final response.
Now let’s move to a real world example involving tools. We will provide web search capability to our agent by defining a custom SerpApi web search tool.
Providers usually have their own built-in tools for web search. However, these tools can be slow or unreliable at times. To get live search data from search engines reliably, we can write a custom tool/function using SerpApi Python SDK.
import json
import os
def google_search(query):
import serpapi
client = serpapi.Client(api_key=os.environ["SERPAPI_KEY"])
results = client.search({
"engine": "google",
"q": query,
})
return [
{
"title": result.get("title"),
"link": result.get("link"),
"snippet": result.get("snippet"),
}
for result in results.get("organic_results", [])[:5]
]
if __name__ == '__main__':
from openai import OpenAI
client = OpenAI(api_key=os.environ["OPENAI_API_KEY"])
tools = [
{
"type": "function",
"name": "google_search",
"description": "Search Google with SerpApi and return web search results.",
"parameters": {
"type": "object",
"properties": {
"query": {
"type": "string",
"description": "The Google search query to run",
}
},
"required": ["query"],
"additionalProperties": False,
},
"strict": True,
}
]
input_list = [
{
"role": "system",
"content": "You are Safar, a travel planner. Use Google search when current destination information would improve your answer.",
},
{
"role": "user",
"content": input("What travel question should I research? "),
},
]
response = client.responses.create(
model="gpt-5.4-mini",
input=input_list,
tools=tools,
tool_choice="required",
)
print("The model responded with:")
print(response.output)
input_list += response.output
for item in response.output:
if item.type != "function_call":
continue
if item.name == "google_search":
args = json.loads(item.arguments)
print(f"The model wants to call google_search with: {args}")
search_results = google_search(args["query"])
print(f"Step 7: SerpApi returned {len(search_results)} search results")
input_list.append({
"type": "function_call_output",
"call_id": item.call_id,
"output": json.dumps(search_results),
})
final_response = client.responses.create(
model="gpt-5.4-mini",
input=input_list,
tools=tools,
)
print(f"Model response: {final_response.output_text}")
Here, we define a google_search() that accepts a query and performs a Google search with the query using SerpApi Python SDK. The function returns the first five search results obtained from Google.
Let’s see the results in action:
What travel question should I research? When is the Tomato festival - La Tomatina happening this year?
The model responded with:
[ResponseFunctionToolCall(arguments='{"query":"La Tomatina 2026 date official"}', call_id='call_mk2KL4xnvR0mexyt2lXFTHgE', name='google_search', type='function_call', id='fc_01af4c5fc07e8479006a192316ab20819bb10273439c89fb9a', namespace=None, status='completed')]
The model wants to call google_search with: {'query': 'La Tomatina 2026 date official'}
Step 7: SerpApi returned 5 search results
Model response: La Tomatina is happening on **Wednesday, August 26, 2026** in **Buñol, Spain**.
If you want, I can also help with:
- tickets
- how to get there from Valencia
- where to stay nearby
This is the core idea behind tool calling. The model does not directly browse the web or fetch data by itself. Instead, it identifies when a tool is needed, asks for that tool to be called, and then uses the returned result to continue the conversation. This separation is useful because the model can focus on reasoning, while tools provide access to external systems and real-time information.
Without the google_search tool, the model would not be able to answer questions that require live data. It should respond with something like: “I don’t have access to real-time information.” By defining the tool, we give the model a safe and structured way to request the information it needs.
MCP
As you build more agents with more tools, a new problem emerges: every tool integration is custom-built and cannot easily be reused elsewhere. If you build a GitHub integration for one agent, you would have to rebuild it from scratch for another. That is where MCP comes in.
Model Context Protocol (MCP) is like USB-C for AI integrations. It is a standard protocol that lets models connect to external tools and data sources in a consistent, reusable way. Instead of building a custom integration for every tool, you write an MCP server once, and any model that supports MCP can use it.
Examples include:
- GitHub MCP
- Figma MCP
- SerpApi MCP
- database MCP servers
- browser MCP servers
- file system MCP servers
With MCP, the model can discover supported functionality and call tools when needed. This makes integrations reusable across different models, clients, and applications. For a small agent, normal tool calling may be enough. For larger systems with many integrations, MCP can make the architecture cleaner.
Let’s see an example of MCP usage in practice. The script below uses the SerpApi MCP server - using this, the agent will be able to call all the SerpApi supported engines like google, google_shopping, amazon, etc.
import os
if __name__ == '__main__':
from openai import OpenAI
client = OpenAI(api_key=os.environ["OPENAI_API_KEY"])
serpapi_mcp_url = f"https://mcp.serpapi.com/{os.environ['SERPAPI_KEY']}/mcp"
response = client.responses.create(
model="gpt-5.4",
tools=[
{
"type": "mcp",
"server_label": "serpapi",
"server_description": "SerpApi MCP server",
"server_url": serpapi_mcp_url,
"require_approval": "never",
}
],
input=[
{
"role": "system",
"content": "You are Cartwise, a shopping assistant. Help users compare products, prices, reviews, and buying options.",
},
{
"role": "user",
"content": input("What do you want to shop for? "),
},
],
)
print("Full model response (includes MCP operations): ")
print(response.output)
print(f"Model response: {response.output_text}")
SerpApi exposes the MCP server via the URL https://mcp.serpapi.com/. Users can supply the API Key via the URL path as seen in the example: https://mcp.serpapi.com/{os.environ['SERPAPI_KEY']}/mcp.
The code here is relatively simpler compared to the tool calling example. We just need to provide the MCP server info via the tools argument:
tools=[
{
"type": "mcp",
"server_label": "serpapi",
"server_description": "SerpApi MCP server",
"server_url": serpapi_mcp_url,
"require_approval": "never",
}
]
From this definition alone, the model can discover supported MCP functionalities and it will be able to autonomously call the MCP server tools based on the user request.
Let’s ask the agent a shopping query. Here, I am asking it to find the price of a mobile device:
What do you want to shop for? Find best price for Moto Razr Ultra phone
Full model response (includes MCP operations):
[
McpListTools(id='mcpl_06176a7178fb9f5c006a17f6c23578819ab2c977e7bc2b0bc7', server_label='serpapi', ...,
McpCall(id='mcp_06176a7178fb9f5c006a17f6c40930819aac6136e6a0f0ced8', arguments='{"params":{"q":"Moto Razr Ultra phone price","engine":"google_shopping","num":10},"mode":"compact"}', name='search', server_label='serpapi', type='mcp_call', approval_request_id=None, error=None, output='{"shopping_results": [{"position": 1, "title": "Motorola Razr Ultra 2025", "product_id": "14521999409488109662", "product_link": ...]}]}', status='completed'),
ResponseOutputMessage(id='msg_06176a7178fb9f5c006a17f6cc3e68819aa688012defa9cf78', content=[ResponseOutputText(annotations=[], text='Best price I found for a **new Moto Razr Ultra** is:...'
]
Model response: Best price I found for a **new Moto Razr Ultra** is:
- **$699.99 at Best Buy** — Motorola Razr Ultra 2025
- was **$1,300**
- rating: **4.0/5** from **520 reviews**
- free delivery by Sat
Also matching:
- **$699.99 at Motorola US** — Motorola Razr 2025
- **$764.00 at Etoren** — Motorola Razr 50 Ultra
- **$1,049.99+** for some Razr 60 Ultra / 2026 variants
The model response includes a series of operations:
- McpListTools: This is the request from the model sent to the MCP server to list available operations. From this call, the model will know that SerpApi has a
google_shoppingengine for shopping searches. - McpCall: Based on the above response, the model calls MCP servers
google_shoppingengine with the queryMoto Razr Ultra phone price. This call will fetch the shopping results via SerpApi MCP. - ResponseOutputMessage: Once the above response is obtained, the model has enough information regarding the prices to formulate its response to the user. The model responds by listing the device price across a number of retailers.
If we omitted the SerpApi MCP definition in the above script, the model should have responded with something like: “I cannot access real-time prices.” This is because the model itself does not have live data access unless we explicitly connect it to external tools or systems. MCP is one way to provide that connection in a standard way.
Now that we have seen how MCP connects agents to external capabilities, let’s look at another way to extend agent behavior: skills.
Skills
While tools handle actions, skills handle behavior. A skill is a reusable set of instructions or a workflow that tells an agent how to perform a specific type of task well.
We have seen tools and MCP which are code-heavy. Tools are code that gets called by the model whereas MCP requires a server implementation according to the Model Context Protocol spec. Skills are relatively simple and can just be a plain text markdown file.
A skill can include:
- steps to follow
- rules
- examples
- output formats
- scripts
- templates
- best practices
Skills are useful for repeated tasks. Examples include writing reports, analyzing PDFs, creating slides, debugging code, or handling customer support. Skills make agents more specialized.
Instead of putting every instruction into the system prompt, we can use skills where the model receives just the skill metadata in the context and will be able to load and use the full skill when the current task needs it.
A skill file is just a markdown file with the below format:
---
name: skill-name
description: A description of what this skill does and when to use it.
---
Skill contents in markdown
Let’s see a real-world example: The SerpApi Search Skill provides instructions for the agent to interact with SerpApi realtime search APIs. You can see the skill.md file, which provides instruction to the model to invoke various SerpApi API calls.
You can see a usage example below, where we use SerpApi skill to build a travel planning agent.
import os
import subprocess
from pathlib import Path
from openai import OpenAI
MODEL = os.getenv("OPENAI_MODEL", "gpt-5.4-mini")
SKILL_PATH = Path(__file__).resolve().parent / "skills" / "serpapi-web-search"
def run_shell_call(shell_call):
print(f"\nModel requested shell call: {shell_call.call_id}")
print(f"Commands: {shell_call.action.commands}")
command_outputs = []
for command in shell_call.action.commands:
print(f"\n[script] Running command: {command}")
result = subprocess.run(
command,
shell=True,
executable="/bin/zsh",
capture_output=True,
text=True,
check=False,
)
print(f"[script] Exit code: {result.returncode}")
if result.stdout:
print(f"[script] stdout:\n{result.stdout[:1500]}")
if result.stderr:
print(f"[script] stderr:\n{result.stderr[:1500]}")
command_outputs.append({
"stdout": result.stdout,
"stderr": result.stderr,
"outcome": {
"type": "exit",
"exit_code": result.returncode,
},
})
output_item = {
"type": "shell_call_output",
"call_id": shell_call.call_id,
"output": command_outputs,
}
if shell_call.action.max_output_length is not None:
output_item["max_output_length"] = shell_call.action.max_output_length
return output_item
if __name__ == "__main__":
client = OpenAI(api_key=os.environ["OPENAI_API_KEY"])
input_list = [
{
"role": "system",
"content": "You are Safar, a travel planning assistant.",
}
]
tools = [
{
"type": "shell",
"environment": {
"type": "local",
"skills": [
{
"name": "serpapi-web-search",
"description": "Search current travel information with the SerpApi CLI.",
"path": str(SKILL_PATH),
}
],
},
}
]
print("Type 'exit' or 'quit' to stop.\n")
waiting_for_user = True
while True:
if waiting_for_user:
user_query = input("You: ")
if user_query.lower() in ["exit", "quit"]:
break
input_list.append({
"role": "user",
"content": user_query,
})
waiting_for_user = False
print("\n[script] Sending request to the model.")
response = client.responses.create(
model=MODEL,
input=input_list,
tools=tools,
)
input_list += response.output
shell_calls = [item for item in response.output if item.type == "shell_call"]
print(f"[script] Shell calls requested: {len(shell_calls)}")
if not shell_calls:
print(f"Model response: {response.output_text}\n")
waiting_for_user = True
continue
for shell_call in shell_calls:
input_list.append(run_shell_call(shell_call))
print("\n[script] Sending shell output back to the model.")
The script uses local skills capability of OpenAI SDK - we have the skill files added in skills/serpapi-web-search folder relative to the scripts parent directory.
Skills can be specified using the below format:
tools = [
{
"type": "shell",
"environment": {
"type": "local",
"skills": [
{
"name": "serpapi-web-search",
"description": "Search current travel information with the SerpApi CLI.",
"path": str(SKILL_PATH),
}
],
},
}
]
We provide the skill name, description and path to the agent. When using skills, OpenAI SDK will emit shell calls that must be run in the terminal. This is needed so that the agent can list and view the full skill file contents that are present locally. We have a run_shell_call() function defined for this. Whenever the model requests for a shell call, we will run this function and pass back the shell results to the model.
Since this example lets the model request shell commands, only run it in a trusted, sandboxed environment. Do not give shell access to untrusted prompts, repositories, or skill files without review.
Now let’s run the agent and ask it a travel planning question. We will ask the model about hotel prices in Goa, India:
Type 'exit' or 'quit' to stop.
You: Find Goa hotel prices for a vacation: two nights from 10 June 26
[script] Sending request to the model.
[script] Shell calls requested: 1
Model requested shell call: call_Cgs0D4tNOZFPZ30GNVJzJNHZ
Commands: ['cd .../skills/serpapi-web-search && cat SKILL.md']
[script] Running command: cd .../skills/serpapi-web-search && cat SKILL.md
[script] Exit code: 0
[script] stdout:
---
name: serpapi-web-search
description: >-
Search the web using SerpApi's 100+ search engines. Use this skill whenever
the user needs current or web-sourced information: ...
[script] Sending shell output back to the model.
[script] Sending request to the model.
[script] Shell calls requested: 1
Model requested shell call: call_Y2PGcBkpSEkzIikqu1H30uRW
Commands: ["cd .../skills/serpapi-web-search && sed -n '1,220p' rules/ENGINES.md"]
[script] Running command: cd .../skills/serpapi-web-search && sed -n '1,220p' rules/ENGINES.md
[script] Exit code: 0
[script] stdout:
# SerpApi Search Engines Catalog
Complete list of 107 SerpApi search engines. Use the `engine` parameter to select the desired search engine.
...
[script] Sending shell output back to the model.
[script] Sending request to the model.
[script] Shell calls requested: 1
Model requested shell call: call_oncTqSWJWQoDbqJS06aJP1OD
Commands: ['cd .../skills/serpapi-web-search && serpapi search engine=google_hotels q="Goa" check_in_date=2026-06-10 check_out_date=2026-06-12 --fields \'properties[0:20]\'']
[script] Running command: cd .../skills/serpapi-web-search && serpapi search engine=google_hotels q="Goa" check_in_date=2026-06-10 check_out_date=2026-06-12 --fields 'properties[0:20]'
[script] Exit code: 0
[script] stdout:
{
"properties": [
{
"type": "hotel",
"name": "Vivanta Goa, Miramar",
"description": "Swanky hotel across from the beach featuring a rooftop pool, a stylish restaurant & a 24-hour gym.",
...
[script] Sending shell output back to the model.
[script] Sending request to the model.
[script] Shell calls requested: 0
Model response: I found live Goa hotel prices for **2 nights: 10 Jun 2026 to 12 Jun 2026**.
Sample prices from Google Hotels:
- **Vivanta Goa, Miramar** — **$82/night**, **$164 total**
- **Aloha Holiday Resort** — **$17/night**, **$33 total**
- Other properties in the Goa search were showing a wide range, from budget stays to luxury resorts.
A few notes:
- These are **current live rates** and can change quickly.
- The prices shown are from hotel search results and may be **before taxes/fees**.
- I searched broadly for **Goa**; if you want, I can narrow it down by:
- **North Goa / South Goa**
- **Budget / mid-range / luxury**
- **Beachfront**
- **2 adults vs family**
If you want, I can make a short list of the **best 10 Goa hotels under a budget you choose**.
As seen from the output, the model initially requested a shell call that runs cat SKILL.md which is to read the skill contents.
With the skill contents obtained, the model proceeds with another shell call sed -n '1,220p' rules/ENGINES.md which lists all SerpApi supported engines. With this data, the model will be able to get all supported SerpApi search engines and choose the best one for the task.
Next, model requests running the command serpapi search engine=google_hotels q="Goa" check_in_date=2026-06-10 check_out_date=2026-06-12 --fields 'properties[0:20]' which uses SerpApi CLI to get results from Google Hotels. We run this shell command on our end and pass the results to the model that includes JSON results from Google Hotels API.
With this data obtained, the model was able to generate its final response and give us suggestions for Hotels to book in Goa along with the prices.
Now that we have seen prompts, memory, tools, MCP, and skills, we can put these pieces into one simple stack.
Agent Capability Stack
An agent can be understood as a stack of capabilities. We have seen the core building blocks of an agent: system prompts, tools, MCP, and skills. Now, let’s compare how they fit together in the agent capability stack.
At the bottom, we have the system prompt. This defines global behavior and constraints.
Then we have skills. Skills provide packaged procedures for specific task types.
Then we have tools. Tools let the agent do things in the world.
Then we have MCP. MCP gives us a standard way to connect models to tools, files, APIs, databases, IDEs, browsers, and other systems.
We can think about the stack like this:
| Layer | Purpose | Use when |
|---|---|---|
| System prompt | Global behavior and constraints | You want rules that apply every turn |
| Skills | Reusable workflows | You want the model to follow a repeatable process |
| Tools | External actions and information | You want the model to call APIs, read files, run code, or fetch live state |
| MCP | Standard integration layer | You want reusable integrations across models and clients |
Use a system prompt for safety boundaries, tone, refusal style, and stable rules.
Use a skill when you want the model to follow a repeatable workflow or use scripts and templates.
Use tools when the model must call external services, fetch live state, create side effects, or interact with the environment.
Use MCP when you want to expose tools and resources through a standard protocol.
Summary and Next Steps
In this tutorial, we started out with the components of an AI agent and built a few simple agents for use cases such as shopping and travel. We provided capabilities to agents using tool calling, MCP, and Skill files.
To explore on your own, you can find the code snippets used in the tutorial in this GitHub repo.
If you are looking for a different SDK or tool to start with like the Claude agent SDK or n8n, we have you covered:
Even though we covered the basics for building simple agents, some important next steps to learn more about are:
- multi-agent systems
- observability
- error handling
- permissions
- context compaction
- evaluation
A multi-agent system has multiple agents, where each agent can be specialized for a specific goal. These agents can communicate with each other. We can also have verifier models that check the output from other models.
Similar to building a backend application, we need observability and error handling for agents. The model can hallucinate, choose the wrong tool, pass bad arguments, or get stuck in a loop. We need a way to monitor this behavior and improve the system over time.
Permissions are also important. An agent that can read files is useful. An agent that can delete files or send emails should be more carefully controlled. We should decide which actions require user approval.
Context compaction is another important idea. As the conversation grows, the agent cannot keep everything forever. It needs to summarize old information and keep only what is useful for the next step.
Evaluation helps us understand whether the agent is actually doing a good job. We can test the agent on sample tasks, check if it used the right tools, verify whether the final answer is correct, and compare outputs across different prompts or models. Without evaluation, it is hard to know if the agent is improving or just producing confident-looking answers.
The best way to understand agents is to build small ones, give them real tasks, inspect their tool calls, and evaluate their outputs. Start with a simple loop, add tools carefully, introduce memory only when needed, and add observability before trusting the agent with important actions. And if your agent needs real-time data access, you can explore SerpApi APIs to extend its capabilities.
Core Dispatch
Core Dispatch #5
Welcome back to Core Dispatch! This edition covers May 18 through June 4, 2026. As promised, Python 3.15.0 beta 2 landed on June 2. Two more milestones are close behind: 3.13.14 and 3.14.6 on June 9, followed by 3.15.0 beta 3 on June 23.
There's also a healthy batch of changes landing for 3.15: an O(n^2) blowup in
unicodedata.normalize() was fixed, the XML parser gained support for
multi-byte encodings, and a round of deprecation warnings went in for the
ast module and abc's abstractclassmethod/abstractstaticmethod/abstractproperty.
On the project side, the Python Security Response Team (PSRT) landed an
initial Python security policy
in the Devguide, giving the vulnerability reporting and response process a
documented home. And dev builds of 3.15+ now report a version like
3.15.0b2+dev instead of the old bare-plus 3.15.0b2+, which
wasn't PEP 440-compliant.
Looking ahead, the EuroPython 2026 Language Summit topics are out, with a lineup spanning a Rust-for-CPython roadmap, the future of free-threading, garbage collection, and the buffer protocol.
If you're interested in CPython internals, Victor Stinner has a great writeup on free threading internals and reference counting that's well worth your time.
As always, if you maintain a package or just like living on the edge, give the latest 3.15 beta a spin and file any issues you find.
Upcoming Releases
- Python 3.13.14 — Jun 09
- Python 3.14.6 — Jun 09
- Python 3.15.0 beta 3 — Jun 23
Official News
- Python 3.15.0 beta 2 is here! — By Hugo van Kemenade
PEP Updates
Merged PRs
- Restore
os.makedirs()ability to applymoderecursively - Add missing ARM64 and RISCV filter in the
lzmamodule - Fix O(n^2) canonical ordering in
unicodedata.normalize() ast: Add deprecation warnings- Raise deprecation warnings for
abc.{abstractclassmethod,abstractstaticmethod,abstractproperty} - Add support of multi-byte encodings in the XML parser
- Improve the PEP 829 batch processing APIs
- Remove
lazy_imports=nonestartup mode - Remove deprecated
'u'type code from thearraymodule - Fix excessive overhead in the Tachyon profiler regarding the cache behavior
devguide: Add an initial Python security policyrelease-tools: Make untagged versions PEP 440-compliant — Untagged/dev builds of CPython now produce PEP 440-compliant version strings.
Discussion
- Understanding PEP discussions — 🆕 🔥 58 new replies · 1.3k views
- PEP 802: Display Syntax for the Empty Set — 🔥 28 new replies · 11.9k views
- PEP 797: Shared Object Proxies — 🔥 14 new replies · 1.9k views
- PEP 828: Supporting 'yield from' in asynchronous generators — 🔥 11 new replies · 4.0k views
- PEP 832: virtual environment discovery — 8 new replies · 4.8k views
- Revisiting PEP 505 – None-aware operators — 4 new replies · 19.9k views
Core Dev Musings
- Free Threading internals: reference counting — By Victor Stinner
Upcoming CFPs & Conferences
- 📋 PyCon Ghana 2026 Deadline — Jun 06
- 📋 PyBeach 2026 Deadline — Jun 08
- GeoPython 2026 — Jun 08
- 📋 PyCon Kenya 2026 Deadline — Jun 09
- 📋 PyCon South Korea 2026 Deadline (extended) — Jun 14
- 📋 Python Ho 2026 Deadline — Jun 15
- PyCon Singapore 2026 — Jun 19
- Python Norte 2026 — Jul 03
- EuroPython 2026 — Jul 13
- SciPy 2026 — Jul 13
Community
- Kirigami — a guided reading workspace for discuss.python.org topics — Early development of a new frontend for DPO that turns a discuss.python.org topic into a guided reading workspace — summary, evidence, participant signals, and the original source posts stay connected.
- EuroPython 2026 Language Summit topics are announced — The talk lineup is out — a Rust-for-CPython roadmap, the future of free-threading, garbage collection, the buffer protocol, the Developer in Residence update, and more.
- An interactive look at CPython's core team over time — A Pyodide-powered chart of the CPython core team — listed members versus those active on python/cpython — with a slider to change the activity window.
One More Thing
""TBC" is "to be confirmed" for Pablo's [Language Summit talk]?"
— Gregory Smith"The Banana Council 🍌"
— Donghee Na
Credits
PyCon Ireland
CFP Deadline Moved to 31 July 2026
We’ve moved the deadline for the PyCon Ireland 2026 Call for Proposals forward to 31 July 2026. It was previously set to 30 August. The submission page on Sessionize already reflects the new date.
If you were planning to submit, please get your proposal in by 31 July 2026.
Why We Brought the Deadline Forward
There are two reasons behind this change, and both are about giving people the time they need to do things well.
Giving the programme committee room to review properly
Building a great schedule takes careful work. With dozens of proposals to read, discuss, and compare, the programme committee needs enough time to give every submission a fair and thorough review. Closing the CFP at the end of August left a tight window between the deadline and the point where we have to lock in the schedule. By moving to 31 July, we give the committee the breathing room to evaluate each proposal on its merits, balance the tracks, and make thoughtful decisions rather than rushed ones.
Giving speakers time to plan their trip to Dublin
PyCon Ireland brings speakers from across Ireland and beyond. Travelling to Dublin means booking flights, arranging accommodation, sorting out time off, and sometimes applying for visas or financial aid. The sooner we can confirm accepted talks, the sooner speakers can start planning, and the less stressful and less expensive that planning tends to be. An earlier deadline means earlier notifications, which is better for everyone making the journey.
What This Means for You
- The new deadline is 31 July 2026. Submit before then.
- Decisions will follow sooner. Bringing the deadline forward lets us notify accepted speakers earlier, so you can lock in travel and accommodation with more lead time.
- Nothing else changes. We still welcome proposals from first-time and experienced speakers alike, across the full range of topics, in both full-talk (30 minutes) and lightning-talk (5 minutes) formats.
Ready to Submit?
Head over to our proposal submission page and tell us what you’d like to talk about. If you have any questions, reach out at contact@python.ie.
We can’t wait to read your proposals, and we’re looking forward to seeing you in Dublin on 17 October 2026.
Bob Belderbos
"Rust Is for People Who Want to Be Punished." Now Jochen Trusts It More Than Python.
Jochen Deister is a lawyer who codes for fun. He has years of Python behind him and no intention of ever being hired to program.
Three months ago, Rust was just a name to him, the language for "the big shots" with a notoriously steep learning curve. Then he built a JSON parser from scratch in Rust, and it ran faster than the equivalent in Python on every dataset he tested, up to 3.5x faster on some. "Holy F" he reacted when he saw the results.
Six weeks of work produced:
- A from-scratch JSON parser, no parsing libraries
- Benchmarks beating Python's standard
jsonmodule (C-accelerated in CPython), up to 3.5x faster - Close to 30 commits in the final week alone, each one a single performance step
- A deliberate 78-error refactor, with the compiler as the guide to a faster implementation
- A new default language: Rust is now the one he reaches for first
Here's how it happened.
The gap
Jochen learned to code on a Commodore VIC-20 with six kilobytes of RAM, then a C64, then a stint in assembly and Turbo Pascal when the bottleneck moved from memory to speed.
Then life took him into law and academia, and he forgot all of it until he picked Python back up years ago.
Python suited him, but it hid the machine. "Python abstracts a lot of these concepts away" he said. "It hides the mechanics".
He'd heard Rust had a notoriously steep learning curve, and he was doing this for fun. "Rust is for people who want to be punished in their life" he figured, and left it there.
The trigger that changed it was small: the last Pybites podcast episode, a $49 lifetime offer on our Rust practice platform, and a remote cabin on the Danish coast where his only job was to keep his kids fed during exam season.
He finished all 61 platform exercises, third on the leaderboard, then shortly after signed up for the cohort for a deeper challenge.
The platform taught him the vocabulary. What it couldn't give him was a real project with a coach reading his code in detail. That's what our cohort is about: six weeks building a JSON parser, one PR review a week with Jim Hodapp, expert Rust coach.
The constraints stopped feeling like constraints
Most people describe their first weeks of Rust as a fight with the borrow checker; the compiler rule that tracks who owns each value and won't let two parts of your code modify the same data at once. Jochen didn't feel it this way at all.
"I never had the feeling that I was fighting the borrow checker. The error messages were my friends right out of the gate. They had a good explanation of the error, but also a hint about what you could do differently."
What hooked him was aesthetics. Run the formatter on a chain of iterator steps and each transformation lands on its own line, readable top to bottom.
"Rust is a beautiful language. It's an aesthetic language. It looks good, and working toward more beautiful code was really something I liked."
That pulled him toward idiomatic Rust on its own. He stopped wanting code that merely worked, the bar he'd accepted in Python, and started wanting code that was safe, performant and idiomatic.
He broke his own code on purpose
Week five, PyO3, was the real step up. PyO3 is the bridge that lets you call a Rust module straight from Python, the same layer Pydantic and Polars are built on. It was the first concept the practice platform hadn't prepared him for, so he leaned on the implementation steps and went slowly.
The clearest sign of how his thinking changed came in the final week. Three of his four benchmark datasets were already beating Python; one wasn't. He suspected the parser was copying the entire input onto the heap instead of borrowing it. So he changed the entry point to take a borrowed string with an explicit lifetime (a lifetime is Rust's way of letting you reference data without copying it, while proving the reference can't outlive the data) and ran cargo check.
It reported 78 errors.
"Those 78 errors were my path of what I needed to fix to get to the results. You change something up the chain and 78 reduces to 50, and so on down the line. It is your implementation guide."
He'd deliberately broken the code, then followed the compiler error by error back to a working, faster version. It's like having a 200% test suite for free; you feel confident making changes.
The rewrite turned a parser that collected every token into a list up front into one that reads tokens on demand in a single pass. Jim's note on the PR: "This is such a clean functional style API for your tokenizer, it's evolved and matured nicely".
The profiler told him where he was wrong
Speed in Rust isn't automatic, and Jochen learned that the hard way. He'd swapped a list for a double-ended queue, proud of it.
"Two days later, when I looked at the profiler, that very line that I was so proud of was now by far the biggest offender."
A profiler measures where a program actually spends its time, so you optimize the real bottleneck instead of a guessed one. His showed the standard-library hash map dominating. He read the docs, realized that map carries protection against denial-of-service attacks he'd never need in a local command-line tool, and replaced it with a stripped-down one. Data-driven, one commit at a time, until the last dataset crossed the line.
Through all of it he kept AI out of the code on purpose. He used it to make himself learn faster, NotebookLM turning Rust docs into podcasts and flashcards, never to write a solution. "Only I write the code" is the rule he gives his AI mentor.
What changed
Ask him how confident he feels starting a new Rust project and he says a 3 out of 10, and means it as a compliment to the language.
"I'm not a total noob anymore. I have a rough understanding of the key concepts, but I also know there's a heck of a lot to learn."
The transfer is in the habits. Rust is now his default for new projects, he caught himself skipping the Python newsletters to read about Rust instead, and the deliberate, idiomatic thinking followed him back into his Python. After years in the Python community, his loyalty quietly shifted:
"I've always liked Python. But it's changed in a way that I think I like Rust more, because of its honesty and because it forces you to think stricter."
His favorite piece of the language is pattern matching, the construct that lets you branch on the shape of a value and pull data out of it in one move. He went deep enough that he used a binding trick his coach hadn't seen before, matching and naming a value in the same arm. Jim's reply on the PR:
"You taught me something I didn't realize Rust has. It's a nice match-and-bind pattern that saves boilerplate code."
The reason he loves it is the same one running through everything he said:
"Computer languages need to be beautiful."
Next up for Jochen: porting a coding agent from Python to Rust, and a privacy tool that strips personal data out of text before it reaches an LLM.
For someone who started three months ago thinking Rust was punishment, that's a real shift. (For more on how Rust rewires the way you write Python, see Learning Rust Made Me a Better Python Developer.)
Here is our full conversation with Jochen about his cohort experience, the parser he built, and the performance work he did:
If you're a Python developer wanting to reach a new level in your career, Rust is a strong contender. Book me in for a call and we'll discuss this further.
June 03, 2026
Real Python
How to Use GitHub Copilot Code Review in Pull Requests
GitHub offers several AI tools under the Copilot umbrella that cover your entire development workflow. Copilot can provide an AI-powered code review shortly after you open a pull request on GitHub. Rather than waiting for a teammate, you can add Copilot as a reviewer to receive context-aware feedback. With access to your entire codebase, it delivers actionable suggestions that you can apply in just a few clicks:
Pull requests are the standard collaborative workflow provided by GitHub and similar services like GitLab to facilitate code review for projects managed with Git. A pull request, or a PR for short, is a formal request to merge code from one branch—or fork—into another, and it’s where code review typically happens.
In practice, code review isn’t always timely or consistent. Some reviewers approve pull requests immediately without much scrutiny, while others leave long lists of minor nitpicks. It can also be difficult to find someone with the right level of experience or enough context about a specific part of the codebase. These issues are common in open-source projects as well, where reviews depend on the limited time of volunteer maintainers.
In this tutorial, you’ll learn how to leverage GitHub Copilot for AI-assisted code review in pull requests and how to integrate it into your workflow to get faster, more structured feedback. Whether you’re working on a commercial project or contributing to an open-source one, Copilot can help you catch issues early and improve your code before it’s merged.
Think of Copilot’s review as a fast first pass. It can reliably flag correctness mistakes and regressions to documented behavior, often before a human reviewer has even opened the PR.
Prerequisites
Before you get started with AI-assisted code reviews, make sure you have the following in place:
- Git and GitHub Knowledge: You should have a basic familiarity with Git and GitHub, including how to create branches, commit changes, and open pull requests.
- Git Client and GitHub CLI: You should have the
gitclient configured in your command line. Additionally, you’ll need the GitHub CLI tool, as it simplifies common GitHub-related tasks. Make sure you’re running v2.88.0 or later, which introduced support for requesting Copilot code reviews from the command line. - GitHub Account: You need a GitHub account with a paid Copilot plan (Pro, Pro+, Business, or Enterprise). To check your subscription status, visit GitHub Copilot settings.
Depending on how you use GitHub, you may already have access to GitHub Copilot through your organization. Sometimes, you may qualify for Copilot under special conditions.
For example, if you’re a student or a teacher, or if you regularly contribute to a popular open-source project, then you might be eligible for free access to GitHub Copilot Pro. Check out GitHub Education to learn more. Keep in mind that GitHub reassesses whether you qualify for free access on a monthly basis.
But even on the free plan, you can still try out Copilot’s code review feature for 30 days at no cost. Just subscribe to GitHub Copilot Pro and cancel before the first billing cycle begins. The trial period is a one-time offer per account, so you won’t be able to start another one after the first one ends.
Note: At the time of writing, GitHub has temporarily paused new paid subscriptions for Copilot due to exceptionally high demand and the associated infrastructure costs. You can read the official announcement on GitHub’s blog to learn more.
To follow along with this tutorial, you’ll also need a GitHub repository where you can freely create branches and pull requests. Although you can create a new repository from scratch or import one from another Git-based hosting service, the quickest option is to download the provided supporting materials. They include a small, hands-on project you’ll be working on:
Get Your Code: Click here to download the free sample code you’ll use to practice AI-assisted code review on a sample FastAPI pull request with GitHub Copilot.
Take the Quiz: Test your knowledge with our interactive “How to Use GitHub Copilot Code Review in Pull Requests” quiz. You’ll receive a score upon completion to help you track your learning progress:
Interactive Quiz
How to Use GitHub Copilot Code Review in Pull RequestsTest your knowledge of GitHub Copilot code review in pull requests, including custom instructions and automatic reviews.
The sample project is a real-time quiz application inspired by Kahoot! and Mentimeter, featuring a FastAPI backend and a mobile-first JavaScript, HTML, and CSS frontend. It allows you to make your own quizzes from scratch—and store them in the human-readable YAML format—or generate a random quiz on the fly using ChatGPT’s API:
Each player is assigned a randomly generated name with an emoji, such as 🐯 Grumpy Tiger, 🦨 Gentle Skunk, or 🐮 Lazy Cow, to keep things light and fun. You can start the server on a local network and have your friends or family connect from their mobile devices using a QR code or a PIN.
Are you ready to dive in?
Step 1: Request a Code Review From GitHub Copilot
If you haven’t already, go ahead and grab the supporting materials. The sample Git repository includes a feature branch with intentional code issues that GitHub Copilot can catch when you request a review. For reference, you’ll also find another branch with the completed code to explore at your own pace:
Get Your Code: Click here to download the free sample code you’ll use to practice AI-assisted code review on a sample FastAPI pull request with GitHub Copilot.
After downloading the materials, upload the local pop-quiz repository—including all branches—to your GitHub account. This will create a remote copy of the repository for your own experimentation. There are several ways to accomplish this. Although you can handle most tasks through the GitHub web interface, the GitHub CLI is often faster and more convenient.
One straightforward approach is to use the GitHub CLI (gh) alongside standard git commands. This allows you to create the repository and push all branches in just two steps once you’re in the downloaded pop-quiz/ directory:
Read the full article at https://realpython.com/github-copilot-code-review/ »
[ Improve Your Python With 🐍 Python Tricks 💌 – Get a short & sweet Python Trick delivered to your inbox every couple of days. >> Click here to learn more and see examples ]
Quiz: How to Use GitHub Copilot Code Review in Pull Requests
In this quiz, you’ll test your understanding of How to Use GitHub Copilot Code Review in Pull Requests.
By working through this quiz, you’ll revisit how to request a review from Copilot on your pull requests, apply or push back on its suggestions, configure automatic reviews, and use custom instructions to make Copilot’s feedback follow your team’s conventions.
[ Improve Your Python With 🐍 Python Tricks 💌 – Get a short & sweet Python Trick delivered to your inbox every couple of days. >> Click here to learn more and see examples ]
Django Weblog
Django security releases issued: 6.0.6 and 5.2.15
In accordance with our security release policy, the Django team is issuing releases for Django 6.0.6 and Django 5.2.15. These releases address the security issues detailed below. We encourage all users of Django to upgrade as soon as possible.
CVE-2026-6873: Signed cookie salt namespace collision in django.http.HttpRequest.get_signed_cookie
get_signed_cookie() derived the signing salt by concatenating the cookie name (key) and salt arguments. When distinct name and salt pairs produced the same concatenation, cookies could be accepted
in a context different from the one where they were signed.
Cookies are now signed with an unambiguous salt derivation. For backwards compatibility, cookies signed by older Django versions are accepted until Django 7.0.
This issue has severity "low" according to the Django security policy.
Thanks to Peng Zhou for the report.
CVE-2026-7666: Potential unencrypted email transmission via STARTTLS in the SMTP backend
When using EMAIL_USE_TLS, a failed STARTTLS handshake could leave a partially-initialized connection that would subsequently be reused for sending email without encryption. This can occur with fail_silently=True, as used by send_mail() and BrokenLinkEmailsMiddleware, among others. Connections configured with EMAIL_USE_SSL are not affected.
This issue has severity "low" according to the Django security policy.
Thanks to Kasper Dupont for the report.
CVE-2026-8404: Potential exposure of private data via case-sensitive Cache-Control directives in UpdateCacheMiddleware
django.middleware.cache.UpdateCacheMiddleware and django.views.decorators.cache.cache_page decorator incorrectly cached responses marked with private Cache-Control directives when using mixed or uppercase values (e.g. Private).
The django.views.decorators.cache.cache_control decorator and django.utils.cache.patch_cache_control() function were not affected, since they normalize directives to lowercase. This issue only affects responses where Cache-Control is set manually.
This issue has severity "low" according to the Django security policy.
Thanks to Ahmed Badawe for the report.
CVE-2026-35193: Potential exposure of private data via missing Vary: Authorization in UpdateCacheMiddleware
django.middleware.cache.UpdateCacheMiddleware and django.views.decorators.cache.cache_page decorator allowed responses to requests bearing an Authorization header (and without Cache-Control: public) to be cached. To conform with the existing mechanism for constructing cache keys, responses to these requests will now vary on Authorization.
This issue has severity "low" according to the Django security policy.
Thanks to Shai Berger for the report.
CVE-2026-48587: Potential exposure of private data via whitespace padding in Vary header
django.middleware.cache.UpdateCacheMiddleware incorrectly cached responses whose Vary header values contained leading or trailing whitespace. Because has_vary_header() failed to strip that whitespace, a response with a Vary: * header (note the trailing space) was not recognized as containing the wildcard, causing it to be stored and potentially served from the cache when it should not have been.
This issue has severity "low" according to the Django security policy.
Thanks to Navid Rezazadeh for the report.
Affected supported versions
- Django main
- Django 6.1 (currently at alpha status)
- Django 6.0
- Django 5.2
Resolution
Patches to resolve the issue have been applied to Django's main, 6.1 (currently at alpha status), 6.0, and 5.2 branches. The patches may be obtained from the following changesets.
CVE-2026-6873: Signed cookie salt namespace collision in django.http.HttpRequest.get_signed_cookie
- On the main branch
- On the 6.1 branch
- On the 6.0 branch
- On the 5.2 branch
CVE-2026-7666: Potential unencrypted email transmission via STARTTLS in the SMTP backend
- On the main branch
- On the 6.1 branch
- On the 6.0 branch
- On the 5.2 branch
CVE-2026-8404: Potential exposure of private data via case-sensitive Cache-Control directives in UpdateCacheMiddleware
- On the main branch
- On the 6.1 branch
- On the 6.0 branch
- On the 5.2 branch
CVE-2026-35193: Potential exposure of private data via missing Vary: Authorization in UpdateCacheMiddleware
- On the main branch
- On the 6.1 branch
- On the 6.0 branch
- On the 5.2 branch
CVE-2026-48587: Potential exposure of private data via whitespace padding in Vary header
- On the main branch
- On the 6.1 branch
- On the 6.0 branch
- On the 5.2 branch
The following releases have been issued
The PGP key ID used for this release is Natalia Bidart: 2EE82A8D9470983E
General notes regarding security reporting
As always, we ask that potential security issues be reported via private email
to security@djangoproject.com, and not via Django's Trac instance, nor via
the Django Forum. Please see
our security policies for further
information.
Python GUIs
Authentication and Authorization with PyQt6 or PySide6 — Secure your desktop applications with login flows, token-based auth, and role-based access control
How can I add authentication and authorization to a PyQt6 application? Is there something built into Qt to make this easier?
When you build a desktop application with PyQt6 or PySide6, sooner or later you'll need to control who can use it and what they can do. Maybe your app connects to a cloud service. Maybe certain features should only be available to administrators. Either way, you need authentication (verifying who the user is) and authorization (deciding what they're allowed to do).
Qt doesn't provide a built-in authentication framework. But that's fine. You can combine Qt's capabilities with Python's networking and security tools to build a solid auth flow for your application.
In this tutorial, we'll walk through the full process: creating a login dialog, authenticating against a remote server, handling tokens, and enabling or disabling parts of your UI based on a user's role.
Approaches to Authentication in Desktop Apps
Before writing any code, it helps to understand the options available when securing a desktop application. The right approach depends on how much security you need and what infrastructure you have.
- Simple login check Your app sends credentials to a remote server at startup. If authentication fails, you disable the UI (partially or entirely). This deters casual users, but a determined hacker could modify the client to bypass the check.
- Token-based unlock After a successful login, the server returns a token or key that unlocks functionality in the app. Without the token, the app can't perform certain operations. This is more secure — the app is genuinely non-functional without a valid token — though once data is decoded into memory, it's theoretically still accessible.
- Server-side execution After authentication, the app sends work to the server, which performs the actual operations. The sensitive logic never runs on the client at all. This is the most secure approach, but it requires server infrastructure to handle the workload.
In the Server-side execution model, the work done on the server doesn't necessarily need to be complex. Transforming or pre-processing some data from one format to another will be enough to deter most attempts at circumvention. However, it's common to to use this technique to hide the algorithmic "secret sauce" completely.
For most applications, the middle ground — authenticating against a remote API and using the returned token to gate access — provides a good balance of security and simplicity. That's what we'll build here.
Your app shouldn't care about the database directly. Instead, it should talk to an API (Application Programming Interface) on your server. The API handles user lookups, password verification, and token generation. Your desktop app just sends HTTP requests and processes the responses.
Setting Up a Simple Auth Server (For Testing)
To test our client application, we need something to authenticate against. We'll create a minimal Flask server that accepts login requests and returns a JSON Web Token (JWT). In a real project, this would be your existing backend, but having a self-contained example makes it easier to experiment.
Install the dependencies for the server:
pip install flask pyjwt
Here's a minimal auth server:
import datetime
import jwt
from flask import Flask, jsonify, request
app = Flask(__name__)
SECRET_KEY = "your-secret-key-change-this"
# In production, use a real database with hashed passwords.
USERS = {
"admin": {"password": "admin123", "role": "admin"},
"viewer": {"password": "viewer123", "role": "viewer"},
}
@app.route("/auth/login", methods=["POST"])
def login():
data = request.get_json()
username = data.get("username", "")
password = data.get("password", "")
user = USERS.get(username)
if user and user["password"] == password:
token = jwt.encode(
{
"username": username,
"role": user["role"],
"exp": datetime.datetime.utcnow()
+ datetime.timedelta(hours=1),
},
SECRET_KEY,
algorithm="HS256",
)
return jsonify(
{"token": token, "role": user["role"], "username": username}
)
return jsonify({"error": "Invalid credentials"}), 401
@app.route("/auth/verify", methods=["GET"])
def verify():
auth_header = request.headers.get("Authorization", "")
if not auth_header.startswith("Bearer "):
return jsonify({"error": "Missing token"}), 401
token = auth_header.split(" ", 1)[1]
try:
payload = jwt.decode(token, SECRET_KEY, algorithms=["HS256"])
return jsonify(
{"username": payload["username"], "role": payload["role"]}
)
except jwt.ExpiredSignatureError:
return jsonify({"error": "Token expired"}), 401
except jwt.InvalidTokenError:
return jsonify({"error": "Invalid token"}), 401
if __name__ == "__main__":
app.run(port=5000, debug=True)
Save this as auth_server.py and run it in a separate terminal:
python auth_server.py
The server exposes two endpoints:
POST /auth/login— accepts a JSON body withusernameandpassword, returns a JWT token.GET /auth/verify— accepts anAuthorization: Bearer <token>header and returns the user info if the token is valid.
This server stores passwords in plain text and uses a hardcoded secret key. In production, you'd hash passwords (using bcrypt or similar) and store the secret key securely. This is purely for demonstration.
Building the Login Dialog
Now let's build the PyQt6 side. We'll start with a login dialog — a modal window where the user enters their credentials. If you're new to dialogs in Qt, see our tutorial on creating dialogs in PyQt6 for a thorough introduction.
Install the client dependencies:
pip install PyQt6 requests
If you're using PySide6, replace
from PyQt6.QtWidgets import ...withfrom PySide6.QtWidgets import ...(and similarly for other Qt modules). The rest of the code is identical.
from PyQt6.QtCore import Qt
from PyQt6.QtWidgets import (
QDialog,
QFormLayout,
QLabel,
QLineEdit,
QPushButton,
QVBoxLayout,
)
class LoginDialog(QDialog):
def __init__(self, parent=None):
super().__init__(parent)
self.setWindowTitle("Login")
self.setFixedSize(350, 200)
layout = QVBoxLayout()
self.form_layout = QFormLayout()
self.username_input = QLineEdit()
self.username_input.setPlaceholderText("Enter your username")
self.form_layout.addRow("Username:", self.username_input)
self.password_input = QLineEdit()
self.password_input.setPlaceholderText("Enter your password")
self.password_input.setEchoMode(QLineEdit.Password)
self.form_layout.addRow("Password:", self.password_input)
layout.addLayout(self.form_layout)
self.login_button = QPushButton("Login")
self.login_button.clicked.connect(self.accept)
layout.addWidget(self.login_button)
self.status_label = QLabel("")
self.status_label.setAlignment(Qt.AlignCenter)
self.status_label.setStyleSheet("color: red;")
layout.addWidget(self.status_label)
self.setLayout(layout)
# Allow pressing Enter to submit.
self.password_input.returnPressed.connect(self.login_button.click)
self.username_input.returnPressed.connect(
self.password_input.setFocus
)
def get_credentials(self):
return (
self.username_input.text().strip(),
self.password_input.text(),
)
def set_status(self, message):
self.status_label.setText(message)
This dialog inherits from QDialog, which gives us the modal behavior we need — when shown with .exec_(), it blocks interaction with the rest of the application until the user either logs in or closes the dialog.
The get_credentials method returns the entered username and password as a tuple. The set_status method lets us display error messages (like "Invalid credentials") directly in the dialog.
Creating an Auth Manager
Rather than scattering authentication logic throughout the application, we'll encapsulate it in a dedicated class. This AuthManager handles login requests, stores the token, and provides the user's role.
import requests
class AuthManager:
def __init__(self, base_url="http://localhost:5000"):
self.base_url = base_url
self.token = None
self.username = None
self.role = None
def login(self, username, password):
"""
Attempt to log in. Returns True on success, False on failure.
Raises an exception on network errors.
"""
response = requests.post(
f"{self.base_url}/auth/login",
json={"username": username, "password": password},
timeout=10,
)
if response.status_code == 200:
data = response.json()
self.token = data["token"]
self.username = data["username"]
self.role = data["role"]
return True
return False
def is_authenticated(self):
return self.token is not None
def get_auth_header(self):
"""Return headers dict with the Bearer token for API requests."""
if self.token:
return {"Authorization": f"Bearer {self.token}"}
return {}
def has_role(self, role):
return self.role == role
def logout(self):
self.token = None
self.username = None
self.role = None
The get_auth_header method is especially useful. Once a user has logged in, you can include this header in any subsequent API call to prove that the request is coming from an authenticated user:
response = requests.get(
"http://localhost:5000/some/protected/endpoint",
headers=auth_manager.get_auth_header(),
timeout=10,
)
Wiring Up the Login Flow
Now we connect the login dialog to the auth manager. The pattern is: show the dialog, grab the credentials, try to authenticate, and either proceed to the main window or show an error.
import sys
from PyQt6.QtWidgets import QApplication, QMessageBox
def attempt_login(auth_manager):
"""
Show the login dialog repeatedly until the user either
successfully authenticates or cancels.
Returns True on successful login, False if cancelled.
"""
dialog = LoginDialog()
while True:
result = dialog.exec_()
if result != QDialog.Accepted:
# User closed the dialog or pressed Cancel.
return False
username, password = dialog.get_credentials()
if not username or not password:
dialog.set_status("Please enter both fields.")
continue
try:
if auth_manager.login(username, password):
return True
else:
dialog.set_status("Invalid username or password.")
except requests.exceptions.ConnectionError:
dialog.set_status("Cannot connect to server.")
except requests.exceptions.Timeout:
dialog.set_status("Connection timed out.")
except requests.exceptions.RequestException as e:
dialog.set_status(f"Error: {e}")
This function keeps showing the login dialog until either the login succeeds or the user dismisses it. Network errors are caught and displayed in the dialog, so the user gets useful feedback without the app crashing.
Building the Main Window with Role-Based Access
The main window of our application will show different features depending on the user's role. Admin users see everything; viewers have a restricted experience. We'll use actions, toolbars, and menus to structure the interface.
from PyQt6.QtWidgets import (
QAction,
QMainWindow,
QMenu,
QMenuBar,
QStatusBar,
QTextEdit,
QToolBar,
)
class MainWindow(QMainWindow):
def __init__(self, auth_manager):
super().__init__()
self.auth_manager = auth_manager
self.setWindowTitle("My Application")
self.setMinimumSize(600, 400)
# Central widget.
self.text_edit = QTextEdit()
self.setCentralWidget(self.text_edit)
# Menu bar.
menu_bar = self.menuBar()
file_menu = menu_bar.addMenu("&File")
self.save_action = QAction("&Save", self)
self.save_action.triggered.connect(self.save_document)
file_menu.addAction(self.save_action)
file_menu.addSeparator()
logout_action = QAction("&Logout", self)
logout_action.triggered.connect(self.handle_logout)
file_menu.addAction(logout_action)
quit_action = QAction("&Quit", self)
quit_action.triggered.connect(self.close)
file_menu.addAction(quit_action)
# Admin-only menu.
self.admin_menu = menu_bar.addMenu("&Admin")
manage_users_action = QAction("&Manage Users", self)
manage_users_action.triggered.connect(self.manage_users)
self.admin_menu.addAction(manage_users_action)
server_settings_action = QAction("&Server Settings", self)
server_settings_action.triggered.connect(self.server_settings)
self.admin_menu.addAction(server_settings_action)
# Status bar.
self.status_bar = QStatusBar()
self.setStatusBar(self.status_bar)
# Apply role-based restrictions.
self.apply_permissions()
def apply_permissions(self):
"""Enable or disable UI elements based on the user's role."""
role = self.auth_manager.role
username = self.auth_manager.username
self.status_bar.showMessage(
f"Logged in as {username} ({role})"
)
if role == "admin":
# Admins get full access.
self.admin_menu.setEnabled(True)
self.save_action.setEnabled(True)
self.text_edit.setReadOnly(False)
elif role == "viewer":
# Viewers can see content but not edit or access admin.
self.admin_menu.setEnabled(False)
self.save_action.setEnabled(False)
self.text_edit.setReadOnly(True)
self.text_edit.setPlaceholderText(
"You have read-only access."
)
else:
# Unknown role: disable everything as a safe default.
self.admin_menu.setEnabled(False)
self.save_action.setEnabled(False)
self.text_edit.setReadOnly(True)
def save_document(self):
QMessageBox.information(
self, "Save", "Document saved (placeholder)."
)
def manage_users(self):
QMessageBox.information(
self, "Admin", "User management (placeholder)."
)
def server_settings(self):
QMessageBox.information(
self, "Admin", "Server settings (placeholder)."
)
def handle_logout(self):
self.auth_manager.logout()
self.close()
The apply_permissions method is where authorization happens. After a successful login, we check the user's role and adjust the UI accordingly. Disabled menu items are grayed out and non-clickable, and the text editor is set to read-only for viewers.
This approach — enabling and disabling widgets based on roles — is the standard pattern for authorization in desktop apps. You can extend it as far as you need: hide entire toolbar sections, show different pages in a stacked widget, or restrict access to specific actions.
Making Authenticated API Requests
Once a user is logged in, you'll often need to make further API calls — fetching data, submitting forms, etc. Each of these requests should include the authentication token so the server can verify the user. For long-running API calls, consider using multithreading with QThreadPool to keep the UI responsive while waiting for server responses.
Here's how you might fetch some protected data:
def fetch_protected_data(auth_manager):
"""Example of making an authenticated API request."""
try:
response = requests.get(
f"{auth_manager.base_url}/auth/verify",
headers=auth_manager.get_auth_header(),
timeout=10,
)
if response.status_code == 200:
return response.json()
elif response.status_code == 401:
# Token expired or invalid — user needs to log in again.
return None
except requests.exceptions.RequestException:
return None
If the server responds with a 401 Unauthorized, that means the token has expired or been revoked. You should handle this gracefully — for example, by showing the login dialog again.
Handling Token Expiration
Tokens expire. When they do, your app needs to respond appropriately rather than silently failing. A common approach is to wrap your API calls in a method that checks for 401 responses and triggers a re-login:
def authenticated_request(auth_manager, method, url, **kwargs):
"""
Make an HTTP request with authentication.
Returns the response, or None if re-authentication fails.
"""
kwargs.setdefault("headers", {})
kwargs["headers"].update(auth_manager.get_auth_header())
kwargs.setdefault("timeout", 10)
try:
response = requests.request(method, url, **kwargs)
if response.status_code == 401:
# Token expired — try to re-authenticate.
if attempt_login(auth_manager):
kwargs["headers"].update(
auth_manager.get_auth_header()
)
response = requests.request(method, url, **kwargs)
else:
return None
return response
except requests.exceptions.RequestException:
return None
This function automatically retries the request with a new token if the first attempt gets a 401. The user sees the login dialog, re-enters their credentials, and the request proceeds as if nothing happened.
To try it out:
- Start the auth server in one terminal:
python auth_server.py - Run the client application in another terminal:
python app.py - Log in as
admin/admin123to see full access, orviewer/viewer123to see restricted access.
Try logging in with the wrong password — the dialog stays open and shows an error. Close the dialog without logging in and the app exits cleanly.
Security Considerations
A few things to keep in mind when implementing auth in a desktop application:
Never store passwords in the client. Your app should only ever send credentials to the server and receive a token back. The token is what you store (in memory, or securely on disk if you want "remember me" functionality).
Use HTTPS in production. Our example uses plain HTTP because it's running locally. In a real deployment, all communication between the client and server should be encrypted with TLS. The requests library handles HTTPS transparently — just change the URL to https://.
Tokens are temporary. JWTs (and most authentication tokens) have an expiration time. Design your app to handle expired tokens gracefully, as shown in the token expiration section above.
Client-side checks are not enough. Disabling a button in the UI doesn't prevent a technically savvy user from calling the underlying function. Any action that matters should be validated on the server side too. The client-side restrictions are a UX convenience, not a security boundary.
Store tokens securely. If you implement a "remember me" feature that persists the token between sessions, use your platform's secure storage — keyring is a good cross-platform Python library for this. Don't write tokens to plain text files. You can also use QSettings to persist non-sensitive user preferences like the last-used username, but avoid storing tokens or credentials there since QSettings does not provide encryption.
For an in-depth guide to building Python GUIs with PyQt6 see my book, Create GUI Applications with Python & Qt6.
Bob Belderbos
How to Tell if Your Python Mock Is Actually Working
A test can pass for the wrong reason. When you're mocking a third-party API call, the test might look green because the real API happened to return an error, not because your mock did anything at all.
This came up in a recent session in our agentic AI cohort where we were looking at a test to verify that converting to an invalid currency raised an exception. The test passed. But something felt off.
The test that passed for the wrong reason
The code under test calls the ExchangeRate API and raises CurrencyConversionError when the response signals failure:
def convert_currency(amount: Decimal, from_currency: str, to_currency: str) -> Decimal:
if from_currency == to_currency:
return amount
response = requests.get(
f"https://v6.exchangerate-api.com/v6/{EXCHANGE_RATE_API_KEY}/pair/{from_currency}/{to_currency}"
)
data = response.json()
if data["result"] != "success":
raise CurrencyConversionError(f"{data['error-type']}")
return Decimal(data["conversion_rate"]) * amountThe test set up a mock_response, patched requests.get to return it (mock_get.return_value = mock_response), but configured it as a successful response:
mock_response.json.return_value = {
"result": "success", # <-- this will never raise CurrencyConversionError
"conversion_rate": 1.5,
}If the mock was intercepting, the function would return normally and pytest.raises would fail. But the test was passing. That meant the mock wasn't intercepting at all: the real API was being hit, and it was returning an error for the bogus "CTM" code.
Proving the mock actually intercepted
My instinct was to add print("calling external api") before requests.get. That proves the code reached that line. It does not prove whether the mock intercepted the call or the real network was hit.
At this point you can put a breakpoint() in the actual requests.get code in your venv, but there is a better way: mock_get.assert_called_once():
with pytest.raises(CurrencyConversionError):
convert_currency(
amount=Decimal("1.00"),
from_currency="CAD",
to_currency="CTM", # Canadian Tire Money, not a real currency
)
mock_get.assert_called_once()If the mock was never called, this assertion fails and tells you directly: your patch didn't intercept the request. If the mock was called, the assertion passes and you know for sure that the test is relying on the mock, not the real API.
Running the test with this assertion in place settled it. Once the patch targeted the right name (the fix in the next section), the mock intercepted the call and pytest.raises failed with DID NOT RAISE. That flip is the proof: a real call for "CTM" would have raised, so a non-raising run means the mock was in control. The earlier green had been the real API answering, never the mock. With the success response still in place, nothing raised. Fixing the response to signal an error made the test pass for the right reason, and assert_called_once() then confirmed the call went through the mock and not the network:
mock_get.return_value.json.return_value = {
"result": "error",
"error-type": "unknown-code",
}Patch where the name is used, not where it's defined
The currency module does import requests then calls requests.get(...), so patching expenses_ai_agent.utils.currency.requests.get targets the call site. With this import requests style, patching requests.get happens to work too, since both names point at the same module object. The rule bites when a module does from requests import get: now get is a local name in the currency module, and you must patch expenses_ai_agent.utils.currency.get, not requests.get. Patching the wrong location is a common mistake that leads to the mock not intercepting and the real API being called.
The cleaned-up test with pytest-mock
Once the mock response was correct and interception was verified, the test got two more improvements. First, the intermediate mock_response variable is unnecessary: chain directly off mock_get.return_value, as in the snippet above. Second, pytest-mock (added with uv add --dev pytest-mock) replaces the nested with patch(...) context managers with a mocker fixture. The result is flatter and easier to scan. Annotated:
def test_bad_currency_conversion_raises(self, mocker):
"""Converting to a non-existing currency should raise an exception."""
# Patch requests.get *as imported inside the currency module* so no
# real HTTP call is made; patch target must match where the name is used
mock_get = mocker.patch("expenses_ai_agent.utils.currency.requests.get")
# Simulate the API response for an unrecognised currency code
mock_get.return_value.json.return_value = {
"result": "error",
"error-type": "unknown-code",
}
with pytest.raises(CurrencyConversionError):
convert_currency(
amount=Decimal("1.00"),
from_currency="CAD",
to_currency="CTM",
)
# Confirm the mock intercepted the call; if this fails, the real API was hit
mock_get.assert_called_once()mocker also handles teardown automatically via the fixture lifecycle, so you don't need with to ensure cleanup.
Another reason to mock: forcing a collision
So far the mock has stood in for a network call. That's not the only reason to reach for one. Here's a test from my simple CRM that stores contacts as files on disk:
def create_contact(
name: str, email: str = "", company: str = "", product: str = ""
) -> str:
contacts_dir().mkdir(parents=True, exist_ok=True)
code = next_code(name)
path = contact_path(code)
if path.exists():
raise FileExistsError(f"Contact {code} already exists")
path.write_text(...)
return codenext_code generates a unique code from the name. To test that creating two contacts with the same code raises FileExistsError, you need both calls to produce the same code. That's nondeterministic by design, so you patch next_code to pin it:
@patch("crm.data.next_code")
def test_cannot_create_contact_with_same_code(mock_next_code):
mock_next_code.return_value = "jd1"
data.create_contact("Jane Doe")
with pytest.raises(FileExistsError):
data.create_contact("Jane Doe")Note the patch target again: crm.data.next_code, where the function is used. Same rule as before. And note that's the only mock here.
Isolation matters as much as the mock, but it doesn't belong in this test. An autouse fixture already points the data dir at a fresh tmp_path:
@pytest.fixture(autouse=True)
def crm_data(tmp_path, monkeypatch):
monkeypatch.setenv("CRM_DATA", str(tmp_path))
(tmp_path / "contacts").mkdir()
return tmp_pathcreate_contact calls path.write_text(...), so the first call writes a real jd1 file. Because every test runs against a fresh tmp_path, that file lives only for the test: the collision can only come from the second call, nothing leaks between runs, and the test fails solely when the duplicate guard fires. Without that isolation, a leftover jd1 from a previous run makes the first call raise, pytest.raises still passes, and you've tested nothing.
Update: I later dropped this mock for an explicit override parameter. Instead of patching next_code, I gave create_contact an optional code parameter (keyword-only, so it can't be passed by accident):
def create_contact(name: str, *, email: str = "", company: str = "",
product: str = "", code: str | None = None) -> str:
...
code = code if code is not None else next_code(name)The test pins the code through the public surface, no patching:
def test_cannot_create_contact_with_same_code():
data.create_contact("Jane Doe")
with pytest.raises(FileExistsError):
data.create_contact("Jane Doe", code="jd1")One naming caveat, since this post points to Harry Percival's "Stop Using Mocks" below: this isn't dependency injection, tempting as it is to call it that. DI would pass next_code itself in and let the test swap a fake. Here I pass the value the dependency would have produced, so it's really an explicit override parameter, the simpler tool. Real DI, with an injected collaborator, comes up at the end of this post.
The trade-off is worth being honest about: I added a production parameter partly to make the test simpler. That's the "test-induced design damage" critics of mocking warn about: a seam that exists only to serve tests. I think it's justified here because code doubles as a real feature: an explicit-code escape hatch for imports or restoring from backup. The test just happens to use it. If the parameter was only added for the test, I'd consider leaving the mock.
Unit vs integration: where does this test belong?
All this then led to a related question:
How should you organize tests that hit real external services?
The convention that holds up in practice:
tests/
├── unit/ # fast, fully mocked, no network, no secrets
└── integration/ # slower, hits real DB / LLM / API endpointsThe currency test above belongs in unit/: it mocks requests.get and never touches the network. A test that actually calls the ExchangeRate API to verify end-to-end behavior belongs in integration/.
A @pytest.mark.integration marker is a lighter-weight way to get the same split without moving files. Register it in pyproject.toml, then skip those tests in CI with pytest -m 'not integration'.
Both work, but the directory structure makes the distinction obvious at a glance. Explicit is better than implicit.
The practical rule: if your test needs an environment variable or some external service to do its real work, it's an integration test. Mock that dependency out and it becomes a unit test. Or put it at the boundary so you can inject a fake in unit tests and the real thing in integration tests (if still needed).
For a practical example of test organization, see this video: Python Unit vs. Functional Testing: Understanding the Difference + Practical Example.
When mocks are the wrong tool
There's a broader point underneath all this. Every time you patch requests.get you're writing a test that's tightly coupled to one import path. Change import requests to from requests import get and every patch breaks. The tests test implementation, not behavior.
I highly recommend watching Harry Percival's PyCon talk "Stop Using Mocks". He makes the case for alternatives: build an adapter class that owns the external call, write a fake in-memory implementation of it, and use dependency injection to pass it in. The repository pattern is the same idea: your test passes in a fake, your production code passes in the real thing, and neither needs patching.
Mocks are still the right choice here: we want to test one small unit whose only external dependency is well contained.
Keep reading
- Two Interesting Scoping Bugs That Made Me Reflect on Object Lifetimes
- The Repository Pattern: Swap Data Sources in One Line
June 02, 2026
PyCoder’s Weekly
Issue #737: Polars 1.41, Email, Great Docs, and More (2026-06-02)
#737 – JUNE 2, 2026
View in Browser »
Announcing Polars 1.41
Polars 1.41 is out and this post covers the new features it includes. Learn about faster parquet metadata decoding, nested subplan elimination, and more.
POLA.RS
Sending Emails With Python
Learn how to send emails with Python using SMTP, attach files, format HTML messages, and personalize bulk emails for your contact list.
REAL PYTHON
Quiz: Sending Emails With Python
Use Python’s standard library to send email through secure SMTP connections, attach files, include HTML content, and route replies.
REAL PYTHON
Your Coding Agent Gets Dumber the Longer It Runs. Here’s the Fix.

Coding agents degrade as context grows. The fix: a multi-role loop where the planner, builder, and reviewer each get isolated context — no stale assumptions, no compounding noise. A practical breakdown from someone who built it. Read the full breakdown
DEPOT sponsor
Great Docs
Talk Python interviews Rich Iannone and Michael Chow from Posit and they talk about a new Python documentation tool called Great Docs.
TALK PYTHON podcast
Articles & Tutorials
Improving Python Through PEPs and Protocols
Have you ever been confused by the naming of modules you’re importing from a package? Is there a standard way to organize and name your Python virtual environments? This week on the show, Brett Cannon returns to discuss the Python Enhancement Proposals (PEPs) he’s been working on recently.
REAL PYTHON podcast
Tame Your Pesky Little Scripts
Over time it is common to accumulate little helper scripts, whether they’re shell scripts, aliases, or custom functions. They are typically tiny things that can become unwieldy to manage. This post shares a few ideas that might help you take back control.
JUHA-MATTI SANTALA
5-Day Live OOP Workshop (Final Chance to Enroll)
The Object-Oriented Python live cohort begins June 8. Five 2-hour sessions Mon to Fri build one growing application end to end, with OOP features introduced as the code starts needing them: classes, the data model, inheritance vs composition, properties, dataclasses.
REAL PYTHON sponsor
Free-Threading vs the GIL in mod_wsgi 6.0.0
Free-threading in mod_wsgi 6.0.0 lets a single process spread Python work across multiple cores. This post is a metrics based comparison between the GIL being enabled and disabled.
GRAHAM DUMPLETON
Notes About Python Email Packages
Chris recently upgraded his personal mail program from Python 2 to Python 3 and this post talks about what needed to change and notes how the newer code works.
CHRIS SIEBENMANN
Learning Path: Perfect Your Python Development Setup
Set up a Python development environment with VS Code, PyCharm, virtual environments, Git, pyenv, Docker, and AI coding tools like Claude Code and Cursor.
REAL PYTHON
Top 7 Python Libraries for Large-Scale Data Processing
This article covers Python libraries that make large-scale data processing faster, more scalable, and easier to manage across modern data workflows.
BALA PRIYA C
Connecting LLMs to Your Data With Python MCP Servers
Build an MCP server in Python that exposes tools, resources, and prompts so AI agents like Cursor can interact with your data.
REAL PYTHON course
How to Make a Scatter Plot in Python With plt.scatter()
Learn how to make scatter plots in Python with plt.scatter() and customize markers by size, color, shape, and transparency.
REAL PYTHON
Two Python Scoping Bugs: A Lesson in Object Lifetimes
Two Python bugs with opposite symptoms but the same root cause: picking the wrong scope for a stateful object.
BOB BELDERBOS
Sentinel Built-In
A quick post about Python 3.15’s new sentinel built-in.
RODRIGO GIRÃO SERRÃO
Projects & Code
Events
Canberra Python Meetup
June 4, 2026
MEETUP.COM
Sydney Python User Group (SyPy)
June 4, 2026
SYPY.ORG
GeoPython 2026
June 8 to June 11, 2026
GEOPYTHON.NET
PiterPy Meetup
June 9, 2026
PITERPY.COM
SciPy 2026, Minneapolis, MN
July 13-19, 2026
SCIPY.ORG • Shared by SciPy Organizers
Happy Pythoning!
This was PyCoder’s Weekly Issue #737.
View in Browser »
[ Subscribe to 🐍 PyCoder’s Weekly 💌 – Get the best Python news, articles, and tutorials delivered to your inbox once a week >> Click here to learn more ]
Real Python
Structuring Your Python Script
You may have begun your Python journey interactively, exploring ideas within Jupyter Notebooks or through the Python REPL. While that’s great for quick experimentation and immediate feedback, you’ll likely find yourself saving code into .py files. However, as your codebase grows, knowing where things should go in your script becomes increasingly important.
Transitioning from interactive environments to structured scripts helps promote readability, enabling better collaboration and more robust development practices. This video course shows you the foundations of organizing a Python script: where the runnable bits go, how to arrange your imports, and how to refactor with constants and a fixed entry point.
By the end of this video course, you’ll know how to:
- Make a script directly executable on Unix-like systems with a shebang line
- Organize your import statements using standard grouping conventions
- Automatically sort imports and format your code with the
rufflinter - Replace hard-coded values with meaningful constants
- Define a clear script entry point using
if __name__ == "__main__"
Without further ado, it’s time to start working through a concrete script and progressively shape it into well-organized, shareable code.
[ Improve Your Python With 🐍 Python Tricks 💌 – Get a short & sweet Python Trick delivered to your inbox every couple of days. >> Click here to learn more and see examples ]













